Targeting phospholipase D in cancer, infection and neurodegenerative disorders
- PMID: 28209987
- PMCID: PMC6040825
- DOI: 10.1038/nrd.2016.252
Targeting phospholipase D in cancer, infection and neurodegenerative disorders
Abstract
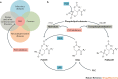
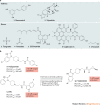
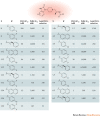
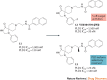
Lipid second messengers have essential roles in cellular function and contribute to the molecular mechanisms that underlie inflammation, malignant transformation, invasiveness, neurodegenerative disorders, and infectious and other pathophysiological processes. The phospholipase D (PLD) isoenzymes PLD1 and PLD2 are one of the major sources of signal-activated phosphatidic acid (PtdOH) generation downstream of a variety of cell-surface receptors, including G protein-coupled receptors (GPCRs), receptor tyrosine kinases (RTKs) and integrins. Recent advances in the development of isoenzyme-selective PLD inhibitors and in molecular genetics have suggested that PLD isoenzymes in mammalian cells and pathogenic organisms may be valuable targets for the treatment of several human diseases. Isoenzyme-selective inhibitors have revealed complex inter-relationships between PtdOH biosynthetic pathways and the role of PtdOH in pathophysiology. PLD enzymes were once thought to be undruggable owing to the ubiquitous nature of PtdOH in cell signalling and concerns that inhibitors would be too toxic for use in humans. However, recent promising discoveries suggest that small-molecule isoenzyme-selective inhibitors may provide novel compounds for a unique approach to the treatment of cancers, neurodegenerative disorders and other afflictions of the central nervous system, and potentially serve as broad-spectrum antiviral and antimicrobial therapeutics.
Conflict of interest statement
H.A.B., C.W.L. and P.G.T. are inventors on patents that protect allosteric phospholipase D inhibitors, as well as a method of treatment using these inhibitors.
Figures


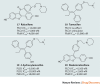
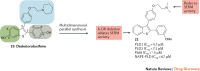
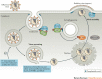
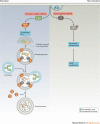
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

