Notch Signaling in Vascular Smooth Muscle Cells
- PMID: 28212801
- PMCID: PMC5964982
- DOI: 10.1016/bs.apha.2016.07.002
Notch Signaling in Vascular Smooth Muscle Cells
Abstract
The Notch signaling pathway is a highly conserved pathway involved in cell fate determination in embryonic development and also functions in the regulation of physiological processes in several systems. It plays an especially important role in vascular development and physiology by influencing angiogenesis, vessel patterning, arterial/venous specification, and vascular smooth muscle biology. Aberrant or dysregulated Notch signaling is the cause of or a contributing factor to many vascular disorders, including inherited vascular diseases, such as cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy, associated with degeneration of the smooth muscle layer in cerebral arteries. Like most signaling pathways, the Notch signaling axis is influenced by complex interactions with mediators of other signaling pathways. This complexity is also compounded by different members of the Notch family having both overlapping and unique functions. Thus, it is vital to fully understand the roles and interactions of each Notch family member in order to effectively and specifically target their exact contributions to vascular disease. In this chapter, we will review the Notch signaling pathway in vascular smooth muscle cells as it relates to vascular development and human disease.
Keywords: Notch signaling; Smooth muscle; Smooth muscle differentiation; Smooth muscle phenotype; Vascular development; Vascular disease.
© 2017 Elsevier Inc. All rights reserved.
Conflict of interest statement
We have no conflicts of interest to report.
Figures

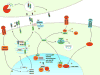
References
-
- Alexander MR, Owens GK. Epigenetic control of smooth muscle cell differentiation and phenotypic switching in vascular development and disease. Annual Review of Physiology. 2012;74:13–40. http://dx.doi.org/10.1146/annurev-physiol-012110-142315. - DOI - PubMed
-
- Andersen P, Uosaki H, Shenje LT, Kwon C. Non-canonical Notch signaling: Emerging role and mechanism. Trends in Cell Biology. 2012;22(5):257–265. http://dx.doi.org/10.1016/j.tcb.2012.02.003. - DOI - PMC - PubMed
-
- Arboleda-Velasquez JF, Primo V, Graham M, James A, Manent J, D’Amore PA. Notch signaling functions in retinal pericyte survival. Investigative Ophthalmology & Visual Science. 2014;55(8):5191–5199. http://dx.doi.org/10.1167/iovs.14-14046. - DOI - PMC - PubMed
-
- Armulik A, Genove G, Betsholtz C. Pericytes: Developmental, physiological, and pathological perspectives, problems, and promises. Developmental Cell. 2011;21(2):193–215. http://dx.doi.org/10.1016/j.devcel.2011.07.001. - DOI - PubMed
-
- Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: Cell fate control and signal integration in development. Science. 1999;284(5415):770–776. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

