Virulence Markers and Phylogenetic Analysis of Escherichia coli Strains with Hybrid EAEC/UPEC Genotypes Recovered from Sporadic Cases of Extraintestinal Infections
- PMID: 28217123
- PMCID: PMC5290387
- DOI: 10.3389/fmicb.2017.00146
Virulence Markers and Phylogenetic Analysis of Escherichia coli Strains with Hybrid EAEC/UPEC Genotypes Recovered from Sporadic Cases of Extraintestinal Infections
Abstract
Virulence genes from different E. coli pathotypes are blended in hybrid strains. E. coli strains with hybrid enteroaggregative/uropathogenic (EAEC/UPEC) genotypes have sporadically emerged causing outbreaks of extraintestinal infections, however their association with routine infections is yet underappreciated. We assessed 258 isolates of E. coli recovered from 86 consecutive cases of extraintestinal infections seeking EAEC and hybrid genotype (EAEC/UPEC) strains. Extensive virulence genotyping was carried out to detect 21 virulence genes, including molecular predictors of EAEC and UPEC strains. Phylogenetic groups and sequence types (STs) were identified, as well as it was performed phylogenetic analyses in order to evaluate whether hybrid EAEC/UPEC strains belonged to intestinal or extraintestinal lineages of E. coli. Adhesion assays were performed to evaluate the biofilm formation by hybrid strains in human urine and cell culture medium (DMEM). Molecular predictors of UPEC were detected in more than 70% of the strains (chuA in 85% and fyuA in 78%). Otherwise, molecular predictors of EAEC (aatA and aggR) were detected in only 3.4% (9/258) of the strains and always along with the UPEC predictor fyuA. Additionally, the pyelonephritis-associated pilus (pap) gene was also detected in all of the hybrid EAEC/UPEC strains. EAEC/UPEC strains were recovered from two cases of community-onset urinary tract infections (UTI) and from a case of bacteremia. Analyses revealed that hybrid EAEC/UPEC strains were phylogenetically positioned in two different clades. Two representative strains, each recovered from UTI and bacteremia, were positioned into a characteristic UPEC clade marked by strains belonging to phylogenetic group D and ST3 (Warwick ST 69). Another hybrid EAEC/UPEC strain was classified as phylogroup A-ST478 and positioned in a commensal clade. Hybrid EAEC/UPEC strains formed biofilms at modest, but perceptible levels either in DMEM or in urine samples. We showed that different lineages of E. coli, at least phylogenetic group A and D, can acquire and gather EAEC and UPEC virulence genes promoting the emergence of hybrid EAEC/UPEC strains.
Keywords: enteroaggregative Escherichia coli; genotyping; hybrid strain; multilocus sequence typing; phylogenetic group; uropathogenic Escherichia coli.
Figures

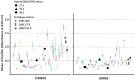
References
-
- Abreu A. G., Fraga T. R., Granados Martínez A. P., Kondo M. Y., Juliano M. A., Juliano L., et al. . (2015). The serine protease pic from enteroaggregative Escherichia coli mediates immune evasion by the direct cleavage of complement proteins. J. Infect. Dis. 212, 106–115. 10.1093/infdis/jiv013 - DOI - PubMed
-
- Aurass P., Prager R., Flieger A. (2011). EHEC/EAEC O104:H4 strain linked with the 2011 German outbreak of haemolytic uremic syndrome enters into the viable but non-culturable state in response to various stresses and resuscitates upon stress relief. Environ. Microbiol. 13, 3139–3148. 10.1111/j.1462-2920.2011.02604.x - DOI - PubMed
-
- Bernier C., Gounon P., Le Bouguénec C. (2002). Identification of an aggregative adhesion fimbria (AAF) type III-encoding operon in enteroaggregative Escherichia coli as a sensitive probe for detecting the AAF-encoding operon family. Infect. Immun. 70, 4302–4311. 10.1128/IAI.70.8.4302-4311.2002 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

