Electrochemistry of the [4Fe4S] Cluster in Base Excision Repair Proteins: Tuning the Redox Potential with DNA
- PMID: 28219007
- PMCID: PMC5423460
- DOI: 10.1021/acs.langmuir.6b04581
Electrochemistry of the [4Fe4S] Cluster in Base Excision Repair Proteins: Tuning the Redox Potential with DNA
Abstract
Escherichia coli endonuclease III (EndoIII) and MutY are DNA glycosylases that contain [4Fe4S] clusters and that serve to maintain the integrity of the genome after oxidative stress. Electrochemical studies on highly oriented pyrolytic graphite (HOPG) revealed that DNA binding by EndoIII leads to a large negative shift in the midpoint potential of the cluster, consistent with stabilization of the oxidized [4Fe4S]3+ form. However, the smooth, hydrophobic HOPG surface is nonideal for working with proteins in the absence of DNA. In this work, we use thin film voltammetry on a pyrolytic graphite edge electrode to overcome these limitations. Improved adsorption leads to substantial signals for both EndoIII and MutY in the absence of DNA, and a large negative potential shift is retained with DNA present. In contrast, the EndoIII mutants E200K, Y205H, and K208E, which provide electrostatic perturbations in the vicinity of the cluster, all show DNA-free potentials within error of wild type; similarly, the presence of negatively charged poly-l-glutamate does not lead to a significant potential shift. Overall, binding to the DNA polyanion is the dominant effect in tuning the redox potential of the [4Fe4S] cluster, helping to explain why all DNA-binding proteins with [4Fe4S] clusters studied to date have similar DNA-bound potentials.
Figures


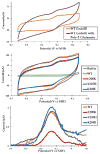
References
-
- Cunningham RP, Asahara H, Bank JF, Scholes CP, Salerno JC, Surerus K, Munck E, McCracken J, Peisach J, Emptage MH. Endonuclease III is an iron-sulfur protein. Biochemistry. 1989;28:4450–4455. - PubMed
-
- Porello SL, Cannon MJ, David SS. A Substrate Recognition Role for the [4Fe4S]2+ Cluster of the DNA Repair Glycosylase MutY. Biochemistry. 1998;37:6465–6475. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

