MYC-driven inhibition of the glutamate-cysteine ligase promotes glutathione depletion in liver cancer
- PMID: 28219903
- PMCID: PMC5376764
- DOI: 10.15252/embr.201643068
MYC-driven inhibition of the glutamate-cysteine ligase promotes glutathione depletion in liver cancer
Abstract
How MYC reprograms metabolism in primary tumors remains poorly understood. Using integrated gene expression and metabolite profiling, we identify six pathways that are coordinately deregulated in primary MYC-driven liver tumors: glutathione metabolism; glycine, serine, and threonine metabolism; aminoacyl-tRNA biosynthesis; cysteine and methionine metabolism; ABC transporters; and mineral absorption. We then focus our attention on glutathione (GSH) and glutathione disulfide (GSSG), as they are markedly decreased in MYC-driven tumors. We find that fewer glutamine-derived carbons are incorporated into GSH in tumor tissue relative to non-tumor tissue. Expression of GCLC, the rate-limiting enzyme of GSH synthesis, is attenuated by the MYC-induced microRNA miR-18a. Inhibition of miR-18a in vivo leads to increased GCLC protein expression and GSH abundance in tumor tissue. Finally, MYC-driven liver tumors exhibit increased sensitivity to acute oxidative stress. In summary, MYC-dependent attenuation of GCLC by miR-18a contributes to GSH depletion in vivo, and low GSH corresponds with increased sensitivity to oxidative stress in tumors. Our results identify new metabolic pathways deregulated in primary MYC tumors and implicate a role for MYC in regulating a major antioxidant pathway downstream of glutamine.
Keywords: MYC; cancer; glutathione; metabolism; miRNA.
© 2017 The Authors.
Figures

Summary of LT2‐MYC conditional transgenic mouse model of MYC‐induced hepatocarcinogenesis. Prolonged MYC overexpression induces tumor nodules that are morphologically and histologically distinct from non‐tumor tissue. MYC protein expression can be turned off in established tumors and correlates with alpha‐fetoprotein (AFP) expression, a marker of aggressive liver cancer (see REG 7 day Western blot). In images, white arrows indicate non‐tumor liver tissue and yellow arrows indicate liver tumor tissue. Scale bars in hematoxylin and eosin‐stained (H&E) sections represent 20 μm.
Transcriptional and biochemical profiling analyses identify six pathways that are significantly altered in LT2‐MYC tumors versus control livers (n = 3 LT2 control and n = 4 LT2‐MYC for transcriptional profiling, n = 7 in each group for biochemical profiling, Fisher's exact test, P < 0.05).
Glutathione pathway (KEGG #ko00480) metabolite abundances segregate LT2‐MYC tumors from control livers by unsupervised hierarchical clustering (n = 7 in each group, LT2 control liver samples in green, LT2‐MYC tumor samples in gray).
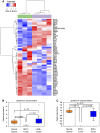
- A
Glutathione pathway transcript expression segregates LT2‐MYC tumors from control livers by unsupervised hierarchical clustering (n = 3 control livers in green, n = 4 tumors in gray).
- B, C
Relative metabolite abundance of GSH (B) or GSSG (C) in tissue samples from murine liver tumors driven by MYC or RAS, as compared to normal liver controls (n = 7 control liver, n = 7 MYC tumor, n = 7 RAS tumor, data represented as box plots with horizontal bar representing the median, box ranges representing the first (bottom) and third (top) quartiles, and vertical bars representing the standard error, unpaired two‐tailed t‐test, Bonferroni adjusted P‐value for multiple comparisons is 0.02).
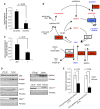
Total glutathione (GSH + GSSG) measured by enzymatic assay in LT2‐MYC tumors versus control livers (n = 5 LT2 control samples, n = 6 LT2‐MYC tumor samples, data represented as mean ± SEM, unpaired two‐tailed t‐test, P = 0.006).
Multiple metabolites and enzymes in the glutathione metabolism pathway are significantly altered in LT2‐MYC tumors versus control livers (unpaired two‐tailed t‐test, P < 0.1). Red = significantly elevated at P < 0.1, blue = significantly depleted at P < 0.05, and red and blue asterisks indicate that individual gamma‐glutamyl amino acids are significantly increased or decreased at P < 0.05. Increased protein expression of the GLS1 isoform of glutaminase was previously reported for LT2‐MYC tumors 11.
Gamma‐glutamylcysteine abundance in MYC‐driven tumors as compared to adjacent non‐tumor tissue (n = 6 each group, data represented as normalized mean ± SEM, paired one‐tailed t‐test, P = 0.04).
Western blot analysis of key enzymes involved in the glutathione metabolism pathway in LT2‐MYC tumors versus non‐tumor LT2 controls (n = 2–3 each as indicated in images, unpaired two‐tailed t‐test on normalized expression, GSS P = 0.7, GLRX5 # P = 0.09, GGT1 *P = 0.05, GSR ***P = 0.0004, G6PDH **P = 0.001, GCLC ***P = 0.0004). For GCLC, LT2‐MYC tumors regressed for 7 days by feeding doxycycline chow are also shown.
Relative incorporation of [U‐13C]‐glutamine into gamma‐glutamylcysteine and GSH in MYC‐driven tumors compared to adjacent non‐tumor liver tissue (n = 6 each group, data represented as normalized mean ± SEM, unpaired two‐tailed t‐test, gamma‐glutamylcysteine P = 0.03, GSH P = 0.004).
Metabolite profiling of gamma‐glutamyl amino acids in LT2‐MYC tumors versus control livers (n = 7 control liver, n = 7 MYC tumor, data represented as normalized mean ± SEM, unpaired two‐tailed t‐test, *P < 0.05, **P < 0.01, ****P < 0.00001).
Metabolite profiling of cysteine–glutathione disulfide and S‐methylglutathione in LT2‐MYC tumors versus control livers (n = 7 control liver, n = 7 MYC tumor, data represented as normalized mean ± SEM, unpaired two‐tailed t‐test, cysteine–glutathione disulfide P = 1.97305E‐07, S‐methylglutathione P = 1.73948E‐09).
Western blot analysis of MYC and GCLC protein expression in MYC‐driven murine liver tumors established by hydrodynamic transfection 28 (n = 4 adjacent non‐tumor (ANT), n = 4 MYC tumor).
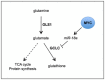
Schematic summary of U‐[13C]‐glutamine tracing experiment in MYC‐driven liver tumor‐bearing mice.
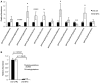
Relative incorporation of [13C]‐glutamine into metabolites in MYC‐driven tumors compared to adjacent non‐tumor liver tissue (n = 6 adjacent non‐tumor (NT), n = 6 MYC tumor (T), all data represented as normalized mean ± SEM, unpaired two‐tailed t‐test, all data significant at P < 0.05).

Quantitative PCR analysis of Gclc mRNA expression in LT2‐MYC tumors, control livers, and liver tumors regressed for 72 h (n = 4 each group, data represented as univariate scatter plot with median, unpaired two‐tailed t‐test, LT2 ctrl versus LT2‐MYC P = 0.0003, LT2‐MYC versus 72 h regression P = 0.0009).
Total glutathione (GSH + GSSG) abundance measured by enzymatic analysis in LT2 control livers, LT2‐MYC tumors, and tumors regressed for 72 h (n = 4 LT2 ctrl, n = 3 LT2‐MYC tumor, n = 3 72 h reg tumor, data represented as univariate scatter plot with median, unpaired one‐tailed t‐test, LT2 ctrl versus LT2‐MYC P = 0.01, LT2‐MYC versus 72 h reg P = 0.14, LT2 ctrl versus 72 h reg P = 0.4, Bonferroni adjusted P‐value is 0.02).
Western blot analysis of HNRNPA1 in LT2‐MYC tumors, control livers, and tumors regressed for 72 h (n = 3 each group).
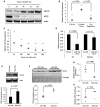
- A
Western blot analysis of GCLC and MYC protein expression in conditional liver tumor cells derived from an LT2‐MYC tumor (Western blot is representative of a minimum of four experimental replicates).
- B
Quantitative PCR (qPCR) analysis of miR‐18a expression in LT2‐MYC tumors, control liver tissues, and tumors regressed for 3 days (n = 4 each group, data represented as univariate scatter plot with median, unpaired two‐tailed t‐test, LT2 ctrl versus LT2‐MYC tumor P = 0.002, LT2‐MYC tumor versus 72 h regression tumors P = 0.001).
- C
qPCR analysis of miR‐18a expression in conditional liver tumor cells treated with doxycycline (data represented as univariate scatter plot with median, data points represent three experimental replicates comprised of three technical replicates each, unpaired two‐tailed t‐test; compared to 0 h: 24 h, P = 0.14; 48 h, P = 0.008; 72 h, P = 0.005; 96 h, P = 0.003, **P‐values for 48 h, 72 h, and 96 h fall below Bonferroni adjusted P‐value of 0.01).
- D
Luciferase reporter expression in cultured murine liver tumor cells treated with a miR‐18a mimic or control. Wt, wild‐type Gclc 3′ UTR; mutated, Gclc 3′ UTR with four base pairs of the putative miR‐18a binding site mutated (data represented as normalized mean ± SEM of three experimental replicates with three technical replicates each, unpaired two‐tailed t‐test, Wt UTR ctrl versus 18a mimic P = 0.0002, mutated UTR control versus 18a mimic P = 0.14).
- E
Western blot (WB) analysis of GCLC protein expression following treatment of cultured LT2‐MYC liver tumor cells with locked nucleic acid (LNA) inhibitors of miR‐18a (WB representative of two experimental replicates with three technical replicates each, pooled data represented as mean ± SEM, unpaired two‐tailed t‐test, P = 0.007).
- F–H
WB of GCLC protein expression (F), qPCR of miR‐18a expression (G), and enzymatic quantitation of GSH abundance (H) in liver tissue samples following treatment of LT2‐MYC tumor‐bearing mice with LNA inhibitors of miR‐18a or control LNA (n = 4 control LNA, n = 3 18a LNA, data represented as univariate scatter plots with median, unpaired two‐tailed t‐test, WB P = 0.002; qPCR P = 0.008; GSH assay P = 0.04).
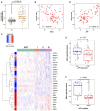
miR‐18a expression in human HCC versus adjacent non‐tumor tissue (n = 96 each group, data represented as box plot with horizontal bar representing the median, box ranges representing the first (bottom) and third (top) quartiles, and vertical bars representing the standard error, Mann–Whitney U‐test, P = 3.58E‐19).
miR‐18a expression inversely correlates with GCLC mRNA expression in human HCC (n = 96, Pearson correlation, Rp = −0.47, two‐tailed t‐test P = 1.315E‐06).
Human HCCs with high miR‐18a expression exhibit a GSH pathway gene expression pattern similar to LT2‐MYC tumors. ANT, adjacent non‐tumor. Roman numerals represent ranked tertiles of increasing miR‐18a expression. Colored bar on left indicates relative gene expression in LT2‐MYC tumors for reference (n = 96 ANT, n = 96 total tumor samples).
miR‐18a expression correlates with AFP mRNA in human HCC (n = 96, Pearson correlation, Rp = 0.70, two‐tailed t‐test P = 2.665E‐15).
Tumor GSH abundance, normalized to matched NT liver tissue, in human HCCs with low (left) versus high (right) serum AFP status (n = 25 each group, data represented as box plot with horizontal bar representing the median, box ranges representing the first (bottom) and third (top) quartiles, and vertical bars representing the standard error, Mann–Whitney U‐test, P = 0.05).
Tumor GSH abundance, normalized to matched NT liver tissue, in hepatic stem cell‐like (HpSC) human HCC versus mature hepatocyte‐like (MH) HCC (n = 15 each group, data represented as box plot with horizontal bar representing the median, box ranges representing the first (bottom) and third (top) quartiles, and vertical bars representing the standard error, Mann–Whitney U‐test, P = 0.004).
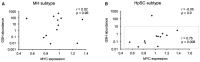
- A, B
Correlation plots of GSH abundance versus MYC mRNA expression in mature hepatocyte (MH) subtype (A) or hepatic stem cell (HpSC) subtype (B) of HCC tumors (samples from 40) (MH: n = 12, Pearson coefficient = 0.02, two‐tailed P = 0.96; HpSC: n = 12, Pearson coefficient = −0.05, two‐tailed P = 0.9; HpSC inset: r = 0.75 and two‐tailed P = 0.008 when an outlier is removed).
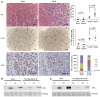
- A
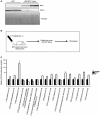
Histological evaluation of MYC‐driven tumors following acute diquat treatment. Upper panel: Representative histology of H&E sections of LT2‐MYC tumors (T) and adjacent non‐tumor (ANT) tissue 24 h after saline (control) or 50 mg/kg diquat treatment. Quantitation is percent area of cell loss per sample (n = 6 saline ANT and T, n = 12 diquat ANT and T, data represented as univariate scatter plot with median, unpaired two‐tailed t‐test, ANT saline versus diquat P = 0.7, T saline versus diquat P = 0.006). Middle panel: representative TUNEL (cell death marker) staining of LT2‐MYC tumors 24 h after saline or 50 mg/kg diquat treatment. Quantitation is average percentage of TUNEL‐positive cells in 10 high‐powered fields for ANT or T samples 24 h after treatment (n = 3 ANT saline, n = 3 ANT diquat, n = 4 T saline, n = 5 T diquat, data represented as univariate scatter plot with median, unpaired two‐tailed t‐test, ANT saline versus diquat P = 0.006, T saline versus diquat P = 0.003). Lower panel: Representative MYC staining of LT2‐MYC tumors 24 h after saline or 50 mg/kg diquat treatment. Quantitation is percent of tumor samples with MYC‐positive score 1+ to 4+ after diquat treatment (n = 6 saline, n = 10 diquat, 6 h and 24 h combined, 1+ corresponds to 1–50% of tumor cells staining positive for MYC, 2+ = 51–74%, 3+ = 75–84%, 4+ = 85–100%, unpaired two‐tailed t‐test, P = 0.02). For all images, scale bar is 50 μm.
- B, C
Western blot analysis of MYC protein expression in LT2‐MYC tumors (T) and adjacent non‐tumor (NT) 6 h (B) and 24 h (C) after 50 mg/kg diquat treatment (n = 2 samples/condition).
References
-
- Osthus RC, Shim H, Kim S, Li Q, Reddy R, Mukherjee M, Xu Y, Wonsey D, Lee LA, Dang CV (2000) Deregulation of glucose transporter 1 and glycolytic gene expression by c‐Myc. J Biol Chem 275: 21797–21800 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

