SAMHD1 enhances nucleoside-analogue efficacy against HIV-1 in myeloid cells
- PMID: 28220857
- PMCID: PMC5318888
- DOI: 10.1038/srep42824
SAMHD1 enhances nucleoside-analogue efficacy against HIV-1 in myeloid cells
Abstract
SAMHD1 is an intracellular enzyme that specifically degrades deoxynucleoside triphosphates into component nucleoside and inorganic triphosphate. In myeloid-derived dendritic cells and macrophages as well as resting T-cells, SAMHD1 blocks HIV-1 infection through this dNTP triphosphohydrolase activity by reducing the cellular dNTP pool to a level that cannot support productive reverse transcription. We now show that, in addition to this direct effect on virus replication, manipulating cellular SAMHD1 activity can significantly enhance or decrease the anti-HIV-1 efficacy of nucleotide analogue reverse transcription inhibitors presumably as a result of modulating dNTP pools that compete for recruitment by viral polymerases. Further, a variety of other nucleotide-based analogues, not normally considered antiretrovirals, such as the anti-herpes drugs Aciclovir and Ganciclovir and the anti-cancer drug Clofarabine are now revealed as potent anti-HIV-1 agents, under conditions of low dNTPs. This in turn suggests novel uses for nucleotide analogues to inhibit HIV-1 in differentiated cells low in dNTPs.
Conflict of interest statement
The authors declare no competing financial interests.
Figures
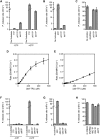
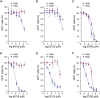
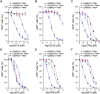
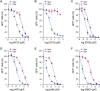
Similar articles
-
A continuous enzyme-coupled assay for triphosphohydrolase activity of HIV-1 restriction factor SAMHD1.Antimicrob Agents Chemother. 2015 Jan;59(1):186-92. doi: 10.1128/AAC.03903-14. Epub 2014 Oct 20. Antimicrob Agents Chemother. 2015. PMID: 25331707 Free PMC article.
-
Anti-HIV host factor SAMHD1 regulates viral sensitivity to nucleoside reverse transcriptase inhibitors via modulation of cellular deoxyribonucleoside triphosphate (dNTP) levels.J Biol Chem. 2013 Jul 12;288(28):20683-91. doi: 10.1074/jbc.M113.472159. Epub 2013 Jun 5. J Biol Chem. 2013. PMID: 23744077 Free PMC article.
-
Dual anti-HIV mechanism of clofarabine.Retrovirology. 2016 Mar 24;13:20. doi: 10.1186/s12977-016-0254-0. Retrovirology. 2016. PMID: 27009333 Free PMC article.
-
The missing link: allostery and catalysis in the anti-viral protein SAMHD1.Biochem Soc Trans. 2019 Aug 30;47(4):1013-1027. doi: 10.1042/BST20180348. Epub 2019 Jul 11. Biochem Soc Trans. 2019. PMID: 31296733 Free PMC article. Review.
-
Intracellular nucleotide levels and the control of retroviral infections.Virology. 2013 Feb 20;436(2):247-54. doi: 10.1016/j.virol.2012.11.010. Epub 2012 Dec 20. Virology. 2013. PMID: 23260109 Free PMC article. Review.
Cited by
-
SAMHD1 as a prognostic and predictive biomarker in stage II colorectal cancer: A multicenter cohort study.Front Oncol. 2022 Aug 1;12:939982. doi: 10.3389/fonc.2022.939982. eCollection 2022. Front Oncol. 2022. PMID: 35978833 Free PMC article.
-
Targeting the DNA damage response and repair in cancer through nucleotide metabolism.Mol Oncol. 2022 Nov;16(21):3792-3810. doi: 10.1002/1878-0261.13227. Epub 2022 May 28. Mol Oncol. 2022. PMID: 35583750 Free PMC article. Review.
-
Positive selection in dNTPase SAMHD1 throughout mammalian evolution.Proc Natl Acad Sci U S A. 2019 Sep 10;116(37):18647-18654. doi: 10.1073/pnas.1908755116. Epub 2019 Aug 26. Proc Natl Acad Sci U S A. 2019. PMID: 31451672 Free PMC article.
-
SAMHD1 Functions and Human Diseases.Viruses. 2020 Mar 31;12(4):382. doi: 10.3390/v12040382. Viruses. 2020. PMID: 32244340 Free PMC article. Review.
-
The Dynamic Interplay between HIV-1, SAMHD1, and the Innate Antiviral Response.Front Immunol. 2017 Nov 10;8:1541. doi: 10.3389/fimmu.2017.01541. eCollection 2017. Front Immunol. 2017. PMID: 29176984 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

