MLL-Rearranged Leukemias-An Update on Science and Clinical Approaches
- PMID: 28232907
- PMCID: PMC5299633
- DOI: 10.3389/fped.2017.00004
MLL-Rearranged Leukemias-An Update on Science and Clinical Approaches
Abstract
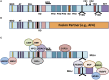
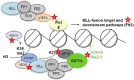
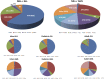
The mixed-lineage leukemia 1 (MLL1) gene (now renamed Lysine [K]-specific MethylTransferase 2A or KMT2A) on chromosome 11q23 is disrupted in a unique group of acute leukemias. More than 80 different partner genes in these fusions have been described, although the majority of leukemias result from MLL1 fusions with one of about six common partner genes. Approximately 10% of all leukemias harbor MLL1 translocations. Of these, two patient populations comprise the majority of cases: patients younger than 1 year of age at diagnosis (primarily acute lymphoblastic leukemias) and young- to-middle-aged adults (primarily acute myeloid leukemias). A much rarer subgroup of patients with MLL1 rearrangements develop leukemia that is attributable to prior treatment with certain chemotherapeutic agents-so-called therapy-related leukemias. In general, outcomes for all of these patients remain poor when compared to patients with non-MLL1 rearranged leukemias. In this review, we will discuss the normal biological roles of MLL1 and its fusion partners, how these roles are hypothesized to be dysregulated in the context of MLL1 rearrangements, and the clinical manifestations of this group of leukemias. We will go on to discuss the progress in clinical management and promising new avenues of research, which may lead to more effective targeted therapies for affected patients.
Keywords: HSCT; chemotherapy; epigenetics; infant leukemia; mixed lineage leukemia; targeted inhibitor.
Figures
References
-
- Butler LH, Slany R, Cui X, Cleary ML, Mason DY. The HRX proto-oncogene product is widely expressed in human tissues and localizes to nuclear structures. Blood (1997) 89(9):3361–70. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

