Epigenome-wide cross-tissue predictive modeling and comparison of cord blood and placental methylation in a birth cohort
- PMID: 28234020
- PMCID: PMC5331917
- DOI: 10.2217/epi-2016-0109
Epigenome-wide cross-tissue predictive modeling and comparison of cord blood and placental methylation in a birth cohort
Abstract
Aim: We compared predictive modeling approaches to estimate placental methylation using cord blood methylation.
Materials & methods: We performed locus-specific methylation prediction using both linear regression and support vector machine models with 174 matched pairs of 450k arrays.
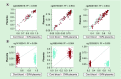
Results: At most CpG sites, both approaches gave poor predictions in spite of a misleading improvement in array-wide correlation. CpG islands and gene promoters, but not enhancers, were the genomic contexts where the correlation between measured and predicted placental methylation levels achieved higher values. We provide a list of 714 sites where both models achieved an R2 ≥0.75.
Conclusion: The present study indicates the need for caution in interpreting cross-tissue predictions. Few methylation sites can be predicted between cord blood and placenta.
Keywords: 450k arrays; DNA methylation; cord blood; epigenetics; methylation prediction; placenta; support vector machine.
Conflict of interest statement
This research was supported by NIH grants R01 HL095606, R01 HL114396, P30 ES023515, R01 ES021357 and R01 NR013945. AC Just was supported by grant R00 ES023450. The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
No writing assistance was utilized in the production of this manuscript.
Figures



Similar articles
-
Locus-specific DNA methylation prediction in cord blood and placenta.Epigenetics. 2019 Apr;14(4):405-420. doi: 10.1080/15592294.2019.1588685. Epub 2019 Mar 18. Epigenetics. 2019. PMID: 30885044 Free PMC article.
-
Early- and late-onset preeclampsia and the DNA methylation of circadian clock and clock-controlled genes in placental and newborn tissues.Chronobiol Int. 2017;34(7):921-932. doi: 10.1080/07420528.2017.1326125. Epub 2017 Jun 14. Chronobiol Int. 2017. PMID: 28613964
-
DNA methylation loci in placenta associated with birthweight and expression of genes relevant for early development and adult diseases.Clin Epigenetics. 2020 Jun 3;12(1):78. doi: 10.1186/s13148-020-00873-x. Clin Epigenetics. 2020. PMID: 32493484 Free PMC article. Clinical Trial.
-
Distinctive aspects of the placental epigenome and theories as to how they arise.Cell Mol Life Sci. 2022 Oct 26;79(11):569. doi: 10.1007/s00018-022-04568-9. Cell Mol Life Sci. 2022. PMID: 36287261 Free PMC article. Review.
-
Epigenetic mechanisms in the placenta related to infant neurodevelopment.Epigenomics. 2018 Mar 1;10(3):321-333. doi: 10.2217/epi-2016-0171. Epub 2018 Jan 30. Epigenomics. 2018. PMID: 29381081 Free PMC article. Review.
Cited by
-
Periconception and Prenatal Exposure to Maternal Perceived Stress and Cord Blood DNA Methylation.Epigenet Insights. 2022 Feb 26;15:25168657221082045. doi: 10.1177/25168657221082045. eCollection 2022. Epigenet Insights. 2022. PMID: 35237744 Free PMC article.
-
Associations between antenatal maternal asthma status and placental DNA methylation.Placenta. 2022 Aug;126:184-195. doi: 10.1016/j.placenta.2022.06.008. Epub 2022 Jul 3. Placenta. 2022. PMID: 35858526 Free PMC article.
-
Epigenetic Alterations of Maternal Tobacco Smoking during Pregnancy: A Narrative Review.Int J Environ Res Public Health. 2021 May 11;18(10):5083. doi: 10.3390/ijerph18105083. Int J Environ Res Public Health. 2021. PMID: 34064931 Free PMC article. Review.
-
Global RNA Expression and DNA Methylation Patterns in Primary Anaplastic Thyroid Cancer.Cancers (Basel). 2020 Mar 13;12(3):680. doi: 10.3390/cancers12030680. Cancers (Basel). 2020. PMID: 32183222 Free PMC article.
-
Cumulative lifetime maternal stress and epigenome-wide placental DNA methylation in the PRISM cohort.Epigenetics. 2018;13(6):665-681. doi: 10.1080/15592294.2018.1497387. Epub 2018 Aug 15. Epigenetics. 2018. PMID: 30001177 Free PMC article.
References
-
- Barrero MJ, Boue S, Izpisua Belmonte JC. Epigenetic mechanisms that regulate cell identity. Cell Stem Cell. 2010;7(5):565–570. - PubMed
-
- Cedar H, Bergman Y. Programming of DNA methylation patterns. Annu. Rev. Biochem. 2012;81:97–117. - PubMed
-
- Kiefer JC. Epigenetics in development. Dev. Dyn. 2007;236(4):1144–1156. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
