Unravelling Endogenous MicroRNA System Dysfunction as a New Pathophysiological Mechanism in Machado-Joseph Disease
- PMID: 28236575
- PMCID: PMC5383579
- DOI: 10.1016/j.ymthe.2017.01.021
Unravelling Endogenous MicroRNA System Dysfunction as a New Pathophysiological Mechanism in Machado-Joseph Disease
Abstract
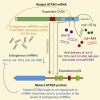
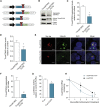
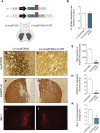
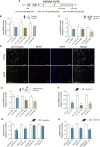
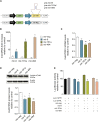
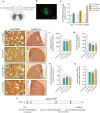
Machado-Joseph disease (MJD) is a genetic neurodegenerative disease caused by an expanded polyglutamine tract within the protein ataxin-3 (ATXN3). Despite current efforts, MJD's mechanism of pathogenesis remains unclear and no disease-modifying treatment is available. Therefore, in this study, we investigated (1) the role of the 3' UTR of ATXN3, a putative microRNA (miRNA) target, (2) whether miRNA biogenesis and machinery are dysfunctional in MJD, and (3) which specific miRNAs target ATXN3-3' UTR and whether they can alleviate MJD neuropathology in vivo. Our results demonstrate that endogenous miRNAs, by targeting sequences in the 3' UTR, robustly reduce ATXN3 expression and aggregation in vitro and neurodegeneration and neuroinflammation in vivo. Importantly, we found an abnormal MJD-associated downregulation of genes involved in miRNA biogenesis and silencing activity. Finally, we identified three miRNAs-mir-9, mir-181a, and mir-494-that interact with the ATXN3-3' UTR and whose expression is dysregulated in human MJD neurons and in other MJD cell and animal models. Furthermore, overexpression of these miRNAs in mice resulted in reduction of mutATXN3 levels, aggregate counts, and neuronal dysfunction. Altogether, these findings indicate that endogenous miRNAs and the 3' UTR of ATXN3 play a crucial role in MJD pathogenesis and provide a promising opportunity for MJD treatment.
Keywords: 3′; Machado-Joseph disease; SCA3; UTR; gene therapy; lentivirus; mechanism of disease; microRNA; microRNA machinery dysregulation.
Copyright © 2017 The American Society of Gene and Cell Therapy. All rights reserved.
Figures

References
-
- Sudarsky L., Coutinho P. Machado-Joseph disease. Clin. Neurosci. 1995;3:17–22. - PubMed
-
- Dürr A., Stevanin G., Cancel G., Duyckaerts C., Abbas N., Didierjean O., Chneiweiss H., Benomar A., Lyon-Caen O., Julien J. Spinocerebellar ataxia 3 and Machado-Joseph disease: clinical, molecular, and neuropathological features. Ann. Neurol. 1996;39:490–499. - PubMed
-
- Kawaguchi Y., Okamoto T., Taniwaki M., Aizawa M., Inoue M., Katayama S., Kawakami H., Nakamura S., Nishimura M., Akiguchi I. CAG expansions in a novel gene for Machado-Joseph disease at chromosome 14q32.1. Nat. Genet. 1994;8:221–228. - PubMed
-
- Seidel K., Siswanto S., Brunt E.R., den Dunnen W., Korf H.W., Rüb U. Brain pathology of spinocerebellar ataxias. Acta Neuropathol. 2012;124:1–21. - PubMed
-
- Paulson H.L., Perez M.K., Trottier Y., Trojanowski J.Q., Subramony S.H., Das S.S., Vig P., Mandel J.L., Fischbeck K.H., Pittman R.N. Intranuclear inclusions of expanded polyglutamine protein in spinocerebellar ataxia type 3. Neuron. 1997;19:333–344. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

