Gene regulatory networking reveals the molecular cue to lysophosphatidic acid-induced metabolic adaptations in ovarian cancer cells
- PMID: 28236660
- PMCID: PMC5527468
- DOI: 10.1002/1878-0261.12046
Gene regulatory networking reveals the molecular cue to lysophosphatidic acid-induced metabolic adaptations in ovarian cancer cells
Abstract
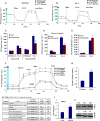
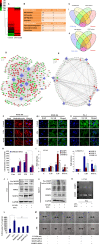
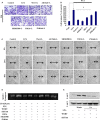
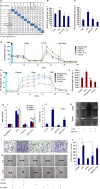
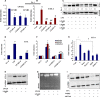
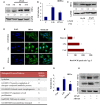
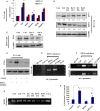
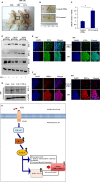
Extravasation and metastatic progression are two main reasons for the high mortality rate associated with cancer. The metastatic potential of cancer cells depends on a plethora of metabolic challenges prevailing within the tumor microenvironment. To achieve higher rates of proliferation, cancer cells reprogram their metabolism, increasing glycolysis and biosynthetic activities. Just why this metabolic reprogramming predisposes cells towards increased oncogenesis remains elusive. The accumulation of myriad oncolipids in the tumor microenvironment has been shown to promote the invasiveness of cancer cells, with lysophosphatidic acid (LPA) being one such critical factor enriched in ovarian cancer patients. Cellular bioenergetic studies confirm that oxidative phosphorylation is suppressed and glycolysis is increased with long exposure to LPA in ovarian cancer cells compared with non-transformed epithelial cells. We sought to uncover the regulatory complexity underlying this oncolipid-induced metabolic perturbation. Gene regulatory networking using RNA-Seq analysis identified the oncogene ETS-1 as a critical mediator of LPA-induced metabolic alterations for the maintenance of invasive phenotype. Moreover, LPA receptor-2 specific PtdIns3K-AKT signaling induces ETS-1 and its target matrix metalloproteases. Abrogation of ETS-1 restores cellular bioenergetics towards increased oxidative phosphorylation and reduced glycolysis, and this effect was reversed by the presence of LPA. Furthermore, the bioenergetic status of LPA-treated ovarian cancer cells mimics hypoxia through induction of hypoxia-inducible factor-1α, which was found to transactivate ets-1. Studies in primary tumors generated in syngeneic mice corroborated the in vitro findings. Thus, our study highlights the phenotypic changes induced by the pro-metastatic factor ETS-1 in ovarian cancer cells. The relationship between enhanced invasiveness and metabolic plasticity further illustrates the critical role of metabolic adaptation of cancer cells as a driver of tumor progression. These findings reveal oncolipid-induced metabolic predisposition as a new mechanism of tumorigenesis and propose metabolic inhibitors as a potential approach for future management of aggressive ovarian cancer.
Keywords: ETS-1; HIF-1α; invasion; lysophosphatidic acid; metabolic adaptations; ovarian cancer.
© 2017 The Authors. Published by FEBS Press and John Wiley & Sons Ltd.
Figures

References
-
- Behrens P, Rothe M, Wellmann A, Krischler J and Wernert N (2001) The Ets‐1 transcription factor is up‐regulated together with MMP 1 and MMP 9 in the stroma of pre‐invasive breast cancer. J Pathol 194, 43–50. - PubMed
-
- Bian D, Mahanivong C, Yu J, Frisch SM, Pan ZK, Ye RD and Huang S (2006) The G12/13‐RhoA signaling pathway contributes to efficient lysophosphatidic acid‐stimulated cell migration. Oncogene 25, 2234–2244. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

