Data on secondary structures and ligand interactions of G-rich oligonucleotides that defy the classical formula for G4 motifs
- PMID: 28243622
- PMCID: PMC5320062
- DOI: 10.1016/j.dib.2017.02.023
Data on secondary structures and ligand interactions of G-rich oligonucleotides that defy the classical formula for G4 motifs
Abstract
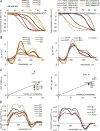
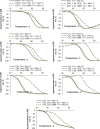
The data provided in this article are related to the research article "The expanding repertoire of G4 DNA structures" [1]. Secondary structures of G-rich oligonucleotides (ONs) that represent "imperfect" G-quadruplex (G4) motifs, i.e., contain truncated or interrupted G-runs, were analyzed by optical methods. Presented data on ON structures include circular dichroism (CD) spectra, thermal difference spectra (TDS) and UV -melting curves of the ONs; and rotational relaxation times (RRT) of ethidium bromide (EtBr) complexes with the ONs. TDS, CD spectra and UV-melting curves can be used to characterize the topologies and thermal stabilities of the ON structures. RRTs are roughly proportional to the hydrodynamic volumes of the complexes and thus can be used to distinguish between inter- and intramolecular ON structures. Presented data on ON interactions with small molecules include fluorescence emission spectra of the G4 sensor thioflavin T (ThT) in complexes with the ONs, and CD-melting curves of the ONs in the presence of G4-stabilizing ligands N-methylmesoporphyrin IX (NMM) and pyridostatin (PDS). These data should be useful for comparative analyses of classical G4s and "defective"G4s, such as quadruplexes with vacancies or bulges.
Keywords: G-quadruplexes; G4 ligands; G4 motifs; Thermal stability.
Figures



Similar articles
-
The expanding repertoire of G4 DNA structures.Biochimie. 2017 Apr;135:54-62. doi: 10.1016/j.biochi.2017.01.003. Epub 2017 Jan 19. Biochimie. 2017. PMID: 28109719
-
Phenanthroline polyazamacrocycles as G-quadruplex DNA binders.Org Biomol Chem. 2018 Apr 18;16(15):2776-2786. doi: 10.1039/c8ob00247a. Org Biomol Chem. 2018. PMID: 29611599
-
Assessment of selectivity of G-quadruplex ligands via an optimised FRET melting assay.Biochimie. 2015 Aug;115:194-202. doi: 10.1016/j.biochi.2015.06.002. Epub 2015 Jun 14. Biochimie. 2015. PMID: 26079222
-
Specific Recognition of Promoter G-Quadruplex DNAs by Small Molecule Ligands and Light-up Probes.ACS Chem Biol. 2019 Oct 18;14(10):2102-2114. doi: 10.1021/acschembio.9b00475. Epub 2019 Oct 4. ACS Chem Biol. 2019. PMID: 31532996 Review.
-
Exploring the Relationship between G-Quadruplex Nucleic Acids and Plants: From Plant G-Quadruplex Function to Phytochemical G4 Ligands with Pharmaceutic Potential.Pharmaceutics. 2022 Nov 4;14(11):2377. doi: 10.3390/pharmaceutics14112377. Pharmaceutics. 2022. PMID: 36365194 Free PMC article. Review.
Cited by
-
Identification and characterization of a G-quadruplex structure in the pre-core promoter region of hepatitis B virus covalently closed circular DNA.J Biol Chem. 2021 Jan-Jun;296:100589. doi: 10.1016/j.jbc.2021.100589. Epub 2021 Mar 24. J Biol Chem. 2021. PMID: 33774051 Free PMC article.
-
Data set on G4 DNA interactions with human proteins.Data Brief. 2018 Mar 9;18:348-359. doi: 10.1016/j.dib.2018.02.081. eCollection 2018 Jun. Data Brief. 2018. PMID: 29896522 Free PMC article.
-
eIF4A alleviates the translational repression mediated by classical secondary structures more than by G-quadruplexes.Nucleic Acids Res. 2018 Apr 6;46(6):3075-3087. doi: 10.1093/nar/gky108. Nucleic Acids Res. 2018. PMID: 29471358 Free PMC article.
References
-
- Varizhuk A., Ischenko D., Tsvetkov V., Novikov R., Kulemin N., Kaluzhny D., Vlasenok M., Naumov V., Smirnov I., Pozmogova G. The expanding repertoire of G4 DNA structures. Biochimie. 2017;135:54–62. - PubMed
-
- Mukundan V.T., Phan A.T. Bulges in G-quadruplexes: broadening the definition of G-quadruplex-forming sequences. J Am. Chem. Soc. 2013;135:5017–5028. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

