Mechanisms of Groucho-mediated repression revealed by genome-wide analysis of Groucho binding and activity
- PMID: 28245789
- PMCID: PMC5331681
- DOI: 10.1186/s12864-017-3589-6
Mechanisms of Groucho-mediated repression revealed by genome-wide analysis of Groucho binding and activity
Abstract
Background: The transcriptional corepressor Groucho (Gro) is required for the function of many developmentally regulated DNA binding repressors, thus helping to define the gene expression profile of each cell during development. The ability of Gro to repress transcription at a distance together with its ability to oligomerize and bind to histones has led to the suggestion that Gro may spread along chromatin. However, much is unknown about the mechanism of Gro-mediated repression and about the dynamics of Gro targeting.
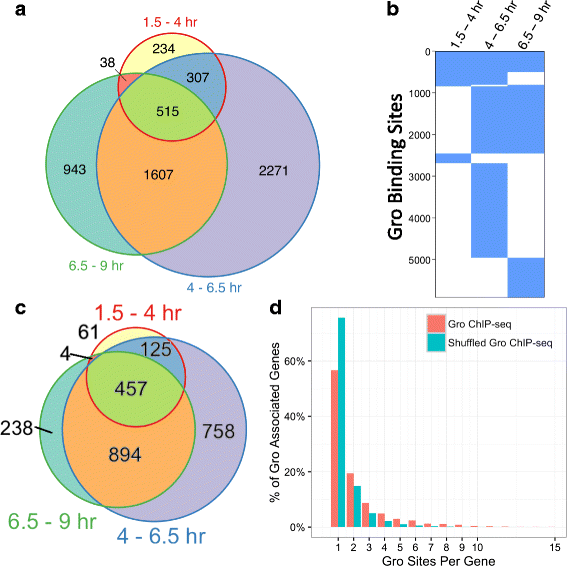
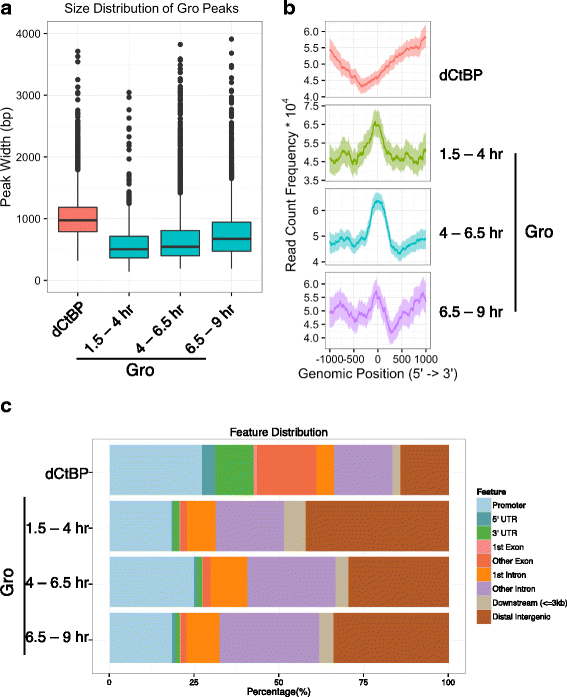
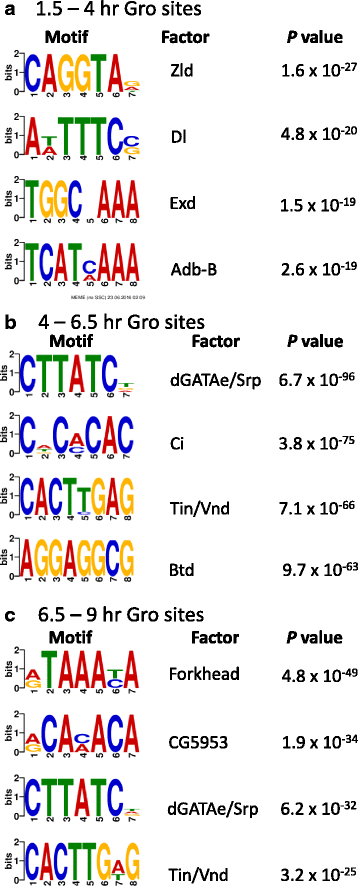
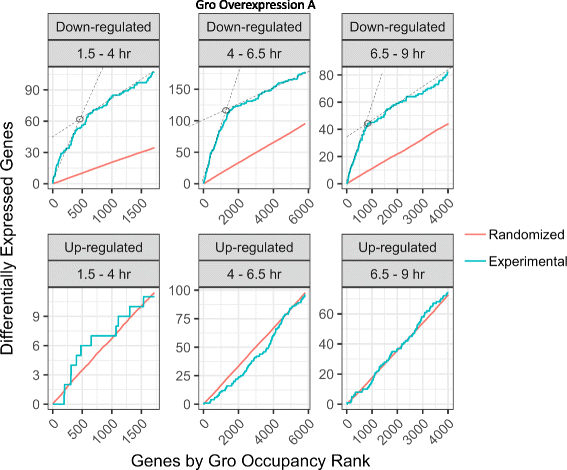
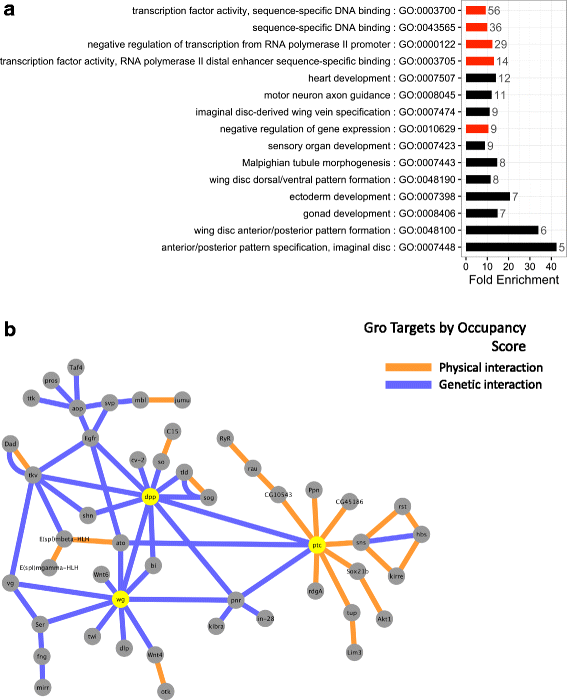
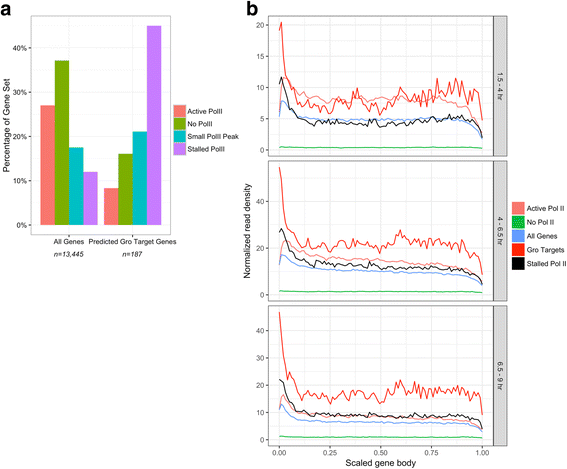
Results: Our chromatin immunoprecipitation sequencing analysis of temporally staged Drosophila embryos shows that Gro binds in a highly dynamic manner primarily to clusters of discrete (<1 kb) segments. Consistent with the idea that Gro may facilitate communication between silencers and promoters, Gro binding is enriched at both cis-regulatory modules, as well as within the promotors of potential target genes. While this Gro-recruitment is required for repression, our data show that it is not sufficient for repression. Integration of Gro binding data with transcriptomic analysis suggests that, contrary to what has been observed for another Gro family member, Drosophila Gro is probably a dedicated repressor. This analysis also allows us to define a set of high confidence Gro repression targets. Using publically available data regarding the physical and genetic interactions between these targets, we are able to place them in the regulatory network controlling development. Through analysis of chromatin associated pre-mRNA levels at these targets, we find that genes regulated by Gro in the embryo are enriched for characteristics of promoter proximal paused RNA polymerase II.
Conclusions: Our findings are inconsistent with a one-dimensional spreading model for long-range repression and suggest that Gro-mediated repression must be regulated at a post-recruitment step. They also show that Gro is likely a dedicated repressor that sits at a prominent highly interconnected regulatory hub in the developmental network. Furthermore, our findings suggest a role for RNA polymerase II pausing in Gro-mediated repression.
Keywords: ChIP-seq; Chromatin-associated RNA-seq, RNA polymerase II pausing; Drosophila embryogenesis; Groucho; RNA-seq; Transcriptional repression.
Figures

Similar articles
-
The Groucho co-repressor is primarily recruited to local target sites in active chromatin to attenuate transcription.PLoS Genet. 2014 Aug 28;10(8):e1004595. doi: 10.1371/journal.pgen.1004595. eCollection 2014 Aug. PLoS Genet. 2014. PMID: 25165826 Free PMC article.
-
The Central Region of the Drosophila Co-repressor Groucho as a Regulatory Hub.J Biol Chem. 2015 Dec 11;290(50):30119-30. doi: 10.1074/jbc.M115.681171. Epub 2015 Oct 19. J Biol Chem. 2015. PMID: 26483546 Free PMC article.
-
A functional interaction between the histone deacetylase Rpd3 and the corepressor groucho in Drosophila development.Genes Dev. 1999 Sep 1;13(17):2218-30. doi: 10.1101/gad.13.17.2218. Genes Dev. 1999. PMID: 10485845 Free PMC article.
-
Groucho/TLE family proteins and transcriptional repression.Gene. 2000 May 16;249(1-2):1-16. doi: 10.1016/s0378-1119(00)00161-x. Gene. 2000. PMID: 10831834 Review.
-
The 'Marx' of Groucho on development and disease.Trends Cell Biol. 2007 Jul;17(7):353-61. doi: 10.1016/j.tcb.2007.07.002. Epub 2007 Jul 20. Trends Cell Biol. 2007. PMID: 17643306 Review.
Cited by
-
Capicua is a fast-acting transcriptional brake.Curr Biol. 2021 Aug 23;31(16):3639-3647.e5. doi: 10.1016/j.cub.2021.05.061. Epub 2021 Jun 23. Curr Biol. 2021. PMID: 34166605 Free PMC article.
-
Loss of TLE3 promotes the mitochondrial program in beige adipocytes and improves glucose metabolism.Genes Dev. 2019 Jul 1;33(13-14):747-762. doi: 10.1101/gad.321059.118. Epub 2019 May 23. Genes Dev. 2019. PMID: 31123067 Free PMC article.
-
Wnt-Dependent Inactivation of the Groucho/TLE Co-repressor by the HECT E3 Ubiquitin Ligase Hyd/UBR5.Mol Cell. 2017 Jul 20;67(2):181-193.e5. doi: 10.1016/j.molcel.2017.06.009. Epub 2017 Jul 6. Mol Cell. 2017. PMID: 28689657 Free PMC article.
-
Normal cell cycle progression requires negative regulation of E2F1 by Groucho during S phase and its relief at G2 phase.Development. 2023 Jun 1;150(11):dev201041. doi: 10.1242/dev.201041. Epub 2023 Jun 1. Development. 2023. PMID: 37260146 Free PMC article.
-
CYCLING DOF FACTOR 1 represses transcription through the TOPLESS co-repressor to control photoperiodic flowering in Arabidopsis.Plant J. 2017 Oct;92(2):244-262. doi: 10.1111/tpj.13649. Epub 2017 Sep 5. Plant J. 2017. PMID: 28752516 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

