Differentiation of ncRNAs from small mRNAs in Escherichia coli O157:H7 EDL933 (EHEC) by combined RNAseq and RIBOseq - ryhB encodes the regulatory RNA RyhB and a peptide, RyhP
- PMID: 28245801
- PMCID: PMC5331693
- DOI: 10.1186/s12864-017-3586-9
Differentiation of ncRNAs from small mRNAs in Escherichia coli O157:H7 EDL933 (EHEC) by combined RNAseq and RIBOseq - ryhB encodes the regulatory RNA RyhB and a peptide, RyhP
Abstract
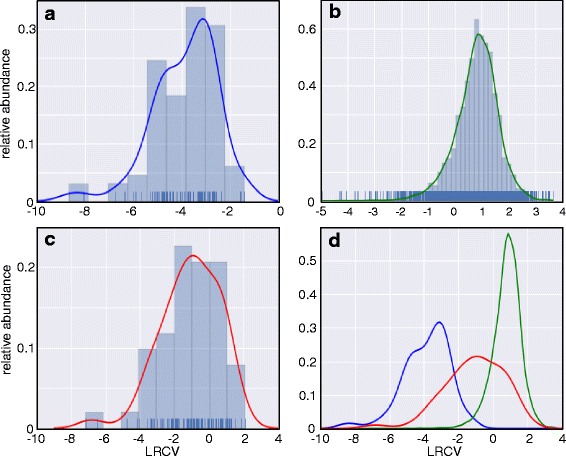
Background: While NGS allows rapid global detection of transcripts, it remains difficult to distinguish ncRNAs from short mRNAs. To detect potentially translated RNAs, we developed an improved protocol for bacterial ribosomal footprinting (RIBOseq). This allowed distinguishing ncRNA from mRNA in EHEC. A high ratio of ribosomal footprints per transcript (ribosomal coverage value, RCV) is expected to indicate a translated RNA, while a low RCV should point to a non-translated RNA.
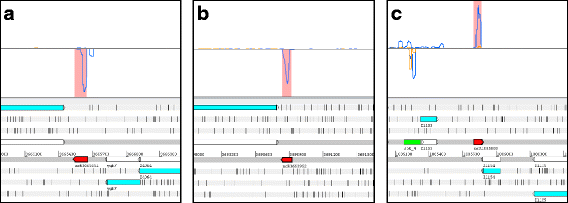
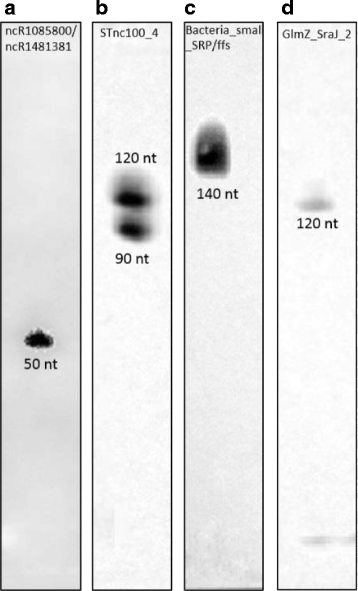
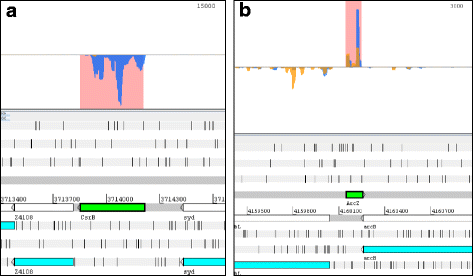
Results: Based on their low RCV, 150 novel non-translated EHEC transcripts were identified as putative ncRNAs, representing both antisense and intergenic transcripts, 74 of which had expressed homologs in E. coli MG1655. Bioinformatics analysis predicted statistically significant target regulons for 15 of the intergenic transcripts; experimental analysis revealed 4-fold or higher differential expression of 46 novel ncRNA in different growth media. Out of 329 annotated EHEC ncRNAs, 52 showed an RCV similar to protein-coding genes, of those, 16 had RIBOseq patterns matching annotated genes in other enterobacteriaceae, and 11 seem to possess a Shine-Dalgarno sequence, suggesting that such ncRNAs may encode small proteins instead of being solely non-coding. To support that the RIBOseq signals are reflecting translation, we tested the ribosomal-footprint covered ORF of ryhB and found a phenotype for the encoded peptide in iron-limiting condition.
Conclusion: Determination of the RCV is a useful approach for a rapid first-step differentiation between bacterial ncRNAs and small mRNAs. Further, many known ncRNAs may encode proteins as well.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

