A Whole-Genome RNA Interference Screen Reveals a Role for Spry2 in Insulin Transcription and the Unfolded Protein Response
- PMID: 28246293
- PMCID: PMC5440024
- DOI: 10.2337/db16-0962
A Whole-Genome RNA Interference Screen Reveals a Role for Spry2 in Insulin Transcription and the Unfolded Protein Response
Abstract
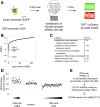
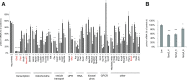
Insulin production by the pancreatic β-cell is required for normal glucose homeostasis. While key transcription factors that bind to the insulin promoter are known, relatively little is known about the upstream regulators of insulin transcription. Using a whole-genome RNA interference screen, we uncovered 26 novel regulators of insulin transcription that regulate diverse processes including oxidative phosphorylation, vesicle traffic, and the unfolded protein response (UPR). We focused on Spry2-a gene implicated in human type 2 diabetes by genome-wide association studies but without a clear connection to glucose homeostasis. We showed that Spry2 is a novel UPR target and its upregulation is dependent on PERK. Knockdown of Spry2 resulted in reduced expression of Serca2, reduced endoplasmic reticulum calcium levels, and induction of the UPR. Spry2 deletion in the adult mouse β-cell caused hyperglycemia and hypoinsulinemia. Our study greatly expands the compendium of insulin promoter regulators and demonstrates a novel β-cell link between Spry2 and human diabetes.
© 2017 by the American Diabetes Association.
Figures




Comment in
-
Checks and Balances-The Limits of β-Cell Endurance to ER Stress.Diabetes. 2017 Jun;66(6):1467-1469. doi: 10.2337/dbi17-0018. Diabetes. 2017. PMID: 28533299 Free PMC article. No abstract available.
References
-
- International Diabetes Federation. IDF Diabetes Atlas, 7th edition [Internet], 2015. Available from http://diabetesatlas.org. Accessed 3 August 2016
-
- Imamura M, Iwata M, Maegawa H, et al. Genetic variants at CDC123/CAMK1D and SPRY2 are associated with susceptibility to type 2 diabetes in the Japanese population. Diabetologia 2011;54:3071–3077 - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

