Evolution of protein-protein interaction networks in yeast
- PMID: 28248977
- PMCID: PMC5382968
- DOI: 10.1371/journal.pone.0171920
Evolution of protein-protein interaction networks in yeast
Abstract
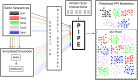
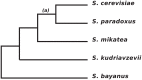
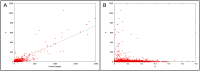
Interest in the evolution of protein-protein and genetic interaction networks has been rising in recent years, but the lack of large-scale high quality comparative datasets has acted as a barrier. Here, we carried out a comparative analysis of computationally predicted protein-protein interaction (PPI) networks from five closely related yeast species. We used the Protein-protein Interaction Prediction Engine (PIPE), which uses a database of known interactions to make sequence-based PPI predictions, to generate high quality predicted interactomes. Simulated proteomes and corresponding PPI networks were used to provide null expectations for the extent and nature of PPI network evolution. We found strong evidence for conservation of PPIs, with lower than expected levels of change in PPIs for about a quarter of the proteome. Furthermore, we found that changes in predicted PPI networks are poorly predicted by sequence divergence. Our analyses identified a number of functional classes experiencing fewer PPI changes than expected, suggestive of purifying selection on PPIs. Our results demonstrate the added benefit of considering predicted PPI networks when studying the evolution of closely related organisms.
Conflict of interest statement
Figures
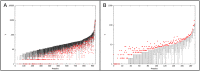
References
-
- Babu M, Krogan NJ, Awrey DE, Emili A, Greenblatt JF. Systematic characterization of the protein interaction network and protein complexes in Saccharomyces cerevisiae using tandem affinity purification and mass spectrometry. Methods in molecular biology (Clifton, NJ). 2009;548:187–207. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

