Combining phylogenetic footprinting with motif models incorporating intra-motif dependencies
- PMID: 28249564
- PMCID: PMC5333389
- DOI: 10.1186/s12859-017-1495-1
Combining phylogenetic footprinting with motif models incorporating intra-motif dependencies
Abstract
Background: Transcriptional gene regulation is a fundamental process in nature, and the experimental and computational investigation of DNA binding motifs and their binding sites is a prerequisite for elucidating this process. Approaches for de-novo motif discovery can be subdivided in phylogenetic footprinting that takes into account phylogenetic dependencies in aligned sequences of more than one species and non-phylogenetic approaches based on sequences from only one species that typically take into account intra-motif dependencies. It has been shown that modeling (i) phylogenetic dependencies as well as (ii) intra-motif dependencies separately improves de-novo motif discovery, but there is no approach capable of modeling both (i) and (ii) simultaneously.
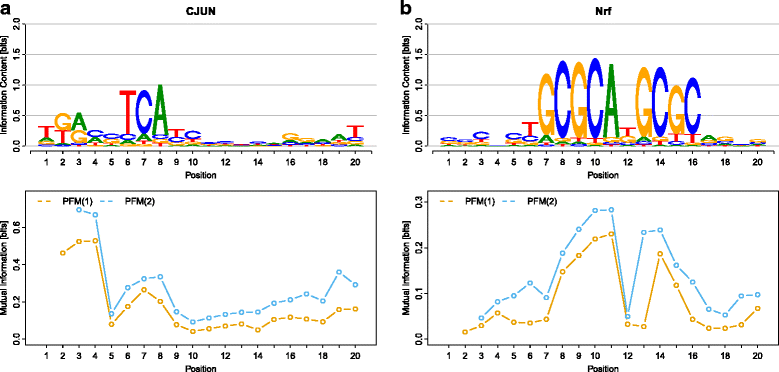
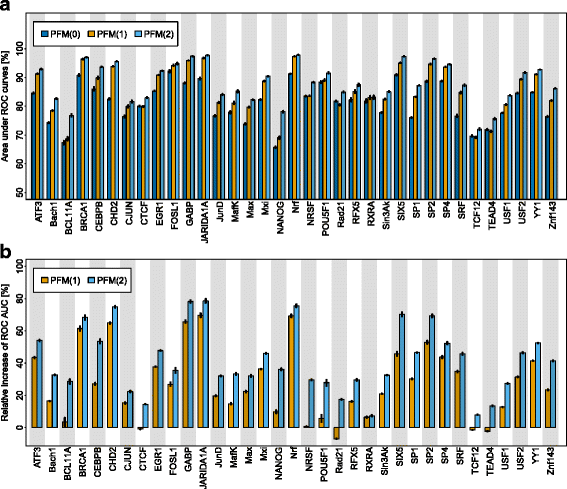
Results: Here, we present an approach for de-novo motif discovery that combines phylogenetic footprinting with motif models capable of taking into account intra-motif dependencies. We study the degree of intra-motif dependencies inferred by this approach from ChIP-seq data of 35 transcription factors. We find that significant intra-motif dependencies of orders 1 and 2 are present in all 35 datasets and that intra-motif dependencies of order 2 are typically stronger than those of order 1. We also find that the presented approach improves the classification performance of phylogenetic footprinting in all 35 datasets and that incorporating intra-motif dependencies of order 2 yields a higher classification performance than incorporating such dependencies of only order 1.
Conclusion: Combining phylogenetic footprinting with motif models incorporating intra-motif dependencies leads to an improved performance in the classification of transcription factor binding sites. This may advance our understanding of transcriptional gene regulation and its evolution.
Keywords: ChIP-Seq; Evolution; Gene regulation; Phylogenetic footprinting; Transcription factor binding sites.
Figures


Similar articles
-
Inferring intra-motif dependencies of DNA binding sites from ChIP-seq data.BMC Bioinformatics. 2015 Nov 9;16:375. doi: 10.1186/s12859-015-0797-4. BMC Bioinformatics. 2015. PMID: 26552868 Free PMC article.
-
Molecular and structural considerations of TF-DNA binding for the generation of biologically meaningful and accurate phylogenetic footprinting analysis: the LysR-type transcriptional regulator family as a study model.BMC Genomics. 2016 Aug 27;17(1):686. doi: 10.1186/s12864-016-3025-3. BMC Genomics. 2016. PMID: 27567672 Free PMC article.
-
Detecting and correcting the binding-affinity bias in ChIP-seq data using inter-species information.BMC Genomics. 2016 May 10;17:347. doi: 10.1186/s12864-016-2682-6. BMC Genomics. 2016. PMID: 27165633 Free PMC article.
-
A survey of DNA motif finding algorithms.BMC Bioinformatics. 2007 Nov 1;8 Suppl 7(Suppl 7):S21. doi: 10.1186/1471-2105-8-S7-S21. BMC Bioinformatics. 2007. PMID: 18047721 Free PMC article. Review.
-
Analysis of Genomic Sequence Motifs for Deciphering Transcription Factor Binding and Transcriptional Regulation in Eukaryotic Cells.Front Genet. 2016 Feb 23;7:24. doi: 10.3389/fgene.2016.00024. eCollection 2016. Front Genet. 2016. PMID: 26941778 Free PMC article. Review.
Cited by
-
Scoring Targets of Transcription in Bacteria Rather than Focusing on Individual Binding Sites.Front Microbiol. 2017 Nov 22;8:2314. doi: 10.3389/fmicb.2017.02314. eCollection 2017. Front Microbiol. 2017. PMID: 29213263 Free PMC article.
-
Unrealistic phylogenetic trees may improve phylogenetic footprinting.Bioinformatics. 2017 Jun 1;33(11):1639-1646. doi: 10.1093/bioinformatics/btx033. Bioinformatics. 2017. PMID: 28130227 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

