A reference gene set for sex pheromone biosynthesis and degradation genes from the diamondback moth, Plutella xylostella, based on genome and transcriptome digital gene expression analyses
- PMID: 28249567
- PMCID: PMC5333385
- DOI: 10.1186/s12864-017-3592-y
A reference gene set for sex pheromone biosynthesis and degradation genes from the diamondback moth, Plutella xylostella, based on genome and transcriptome digital gene expression analyses
Abstract
Background: Female moths synthesize species-specific sex pheromone components and release them to attract male moths, which depend on precise sex pheromone chemosensory system to locate females. Two types of genes involved in the sex pheromone biosynthesis and degradation pathways play essential roles in this important moth behavior. To understand the function of genes in the sex pheromone pathway, this study investigated the genome-wide and digital gene expression of sex pheromone biosynthesis and degradation genes in various adult tissues in the diamondback moth (DBM), Plutella xylostella, which is a notorious vegetable pest worldwide.
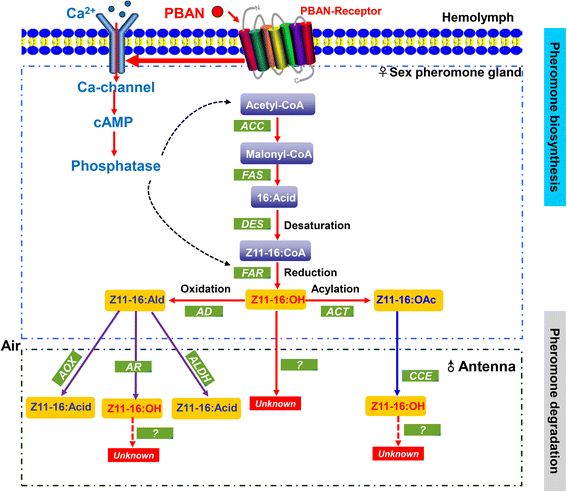
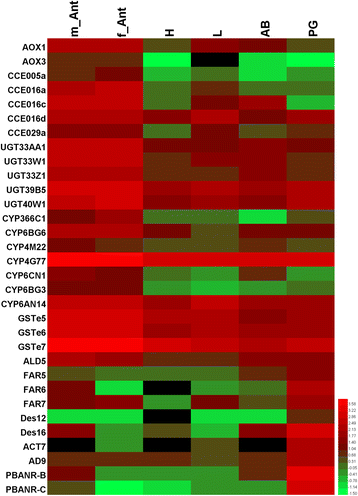
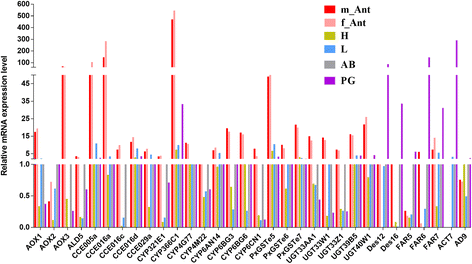
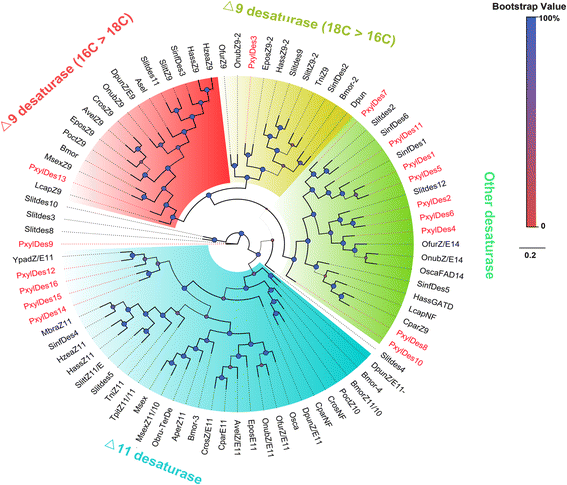
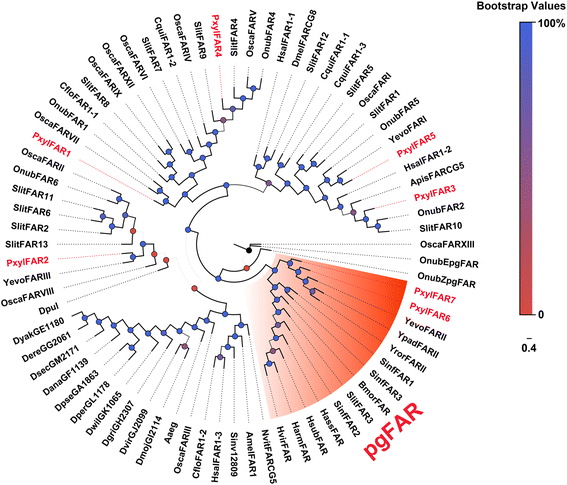
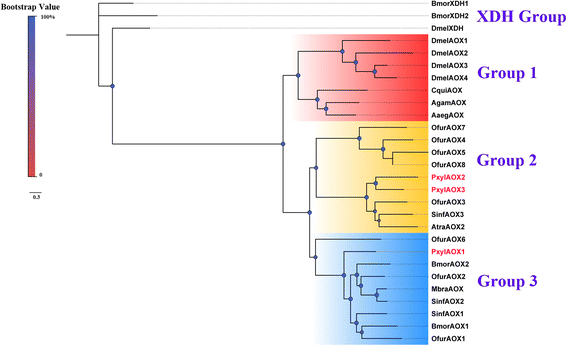
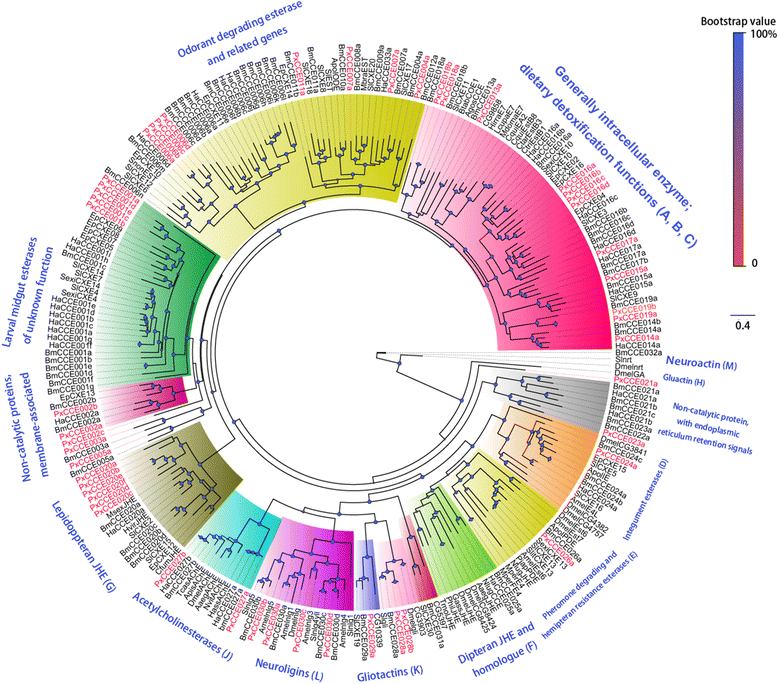
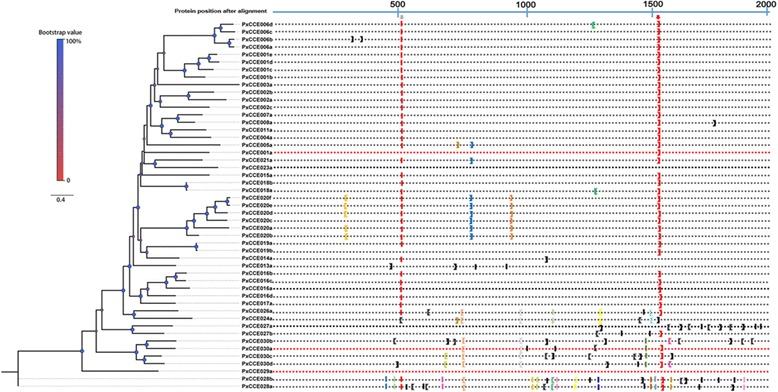
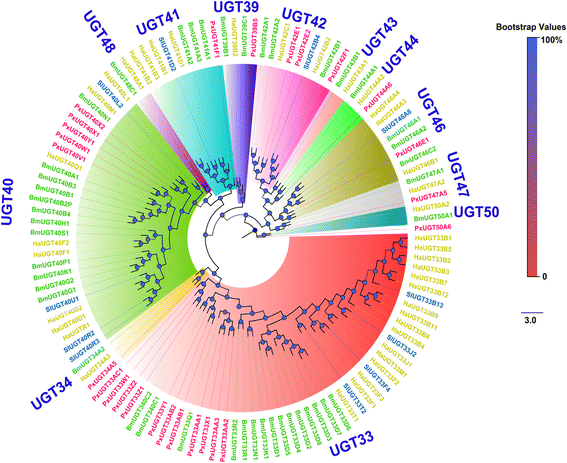
Results: A massive transcriptome data (at least 39.04 Gb) was generated by sequencing 6 adult tissues including male antennae, female antennae, heads, legs, abdomen and female pheromone glands from DBM by using Illumina 4000 next-generation sequencing and mapping to a published DBM genome. Bioinformatics analysis yielded a total of 89,332 unigenes among which 87 transcripts were putatively related to seven gene families in the sex pheromone biosynthesis pathway. Among these, seven [two desaturases (DES), three fatty acyl-CoA reductases (FAR) one acetyltransferase (ACT) and one alcohol dehydrogenase (AD)] were mainly expressed in the pheromone glands with likely function in the three essential sex pheromone biosynthesis steps: desaturation, reduction, and esterification. We also identified 210 odorant-degradation related genes (including sex pheromone-degradation related genes) from seven major enzyme groups. Among these genes, 100 genes are new identified and two aldehyde oxidases (AOXs), one aldehyde dehydrogenase (ALDH), five carboxyl/cholinesterases (CCEs), five UDP-glycosyltransferases (UGTs), eight cytochrome P450 (CYP) and three glutathione S-transferases (GSTs) displayed more robust expression in the antennae, and thus are proposed to participate in the degradation of sex pheromone components and plant volatiles.
Conclusions: To date, this is the most comprehensive gene data set of sex pheromone biosynthesis and degradation enzyme related genes in DBM created by genome- and transcriptome-wide identification, characterization and expression profiling. Our findings provide a basis to better understand the function of genes with tissue enriched expression. The results also provide information on the genes involved in sex pheromone biosynthesis and degradation, and may be useful to identify potential gene targets for pest control strategies by disrupting the insect-insect communication using pheromone-based behavioral antagonists.
Keywords: Aldehyde oxidase; Carboxyl/Cholinesterase; Desaturase; Detoxification; Fatty acyl reductase; Pheromone-biosynthesis enzymes; Pheromone-degrading enzymes.
Figures
Similar articles
-
Identification of genes expressed in the sex pheromone gland of the black cutworm Agrotis ipsilon with putative roles in sex pheromone biosynthesis and transport.BMC Genomics. 2013 Sep 22;14:636. doi: 10.1186/1471-2164-14-636. BMC Genomics. 2013. PMID: 24053512 Free PMC article.
-
Identification of the pheromone biosynthesis genes from the sex pheromone gland transcriptome of the diamondback moth, Plutella xylostella.Sci Rep. 2017 Nov 24;7(1):16255. doi: 10.1038/s41598-017-16518-8. Sci Rep. 2017. PMID: 29176628 Free PMC article.
-
Identification of putative Type-I sex pheromone biosynthesis-related genes expressed in the female pheromone gland of Streltzoviella insularis.PLoS One. 2020 Jan 16;15(1):e0227666. doi: 10.1371/journal.pone.0227666. eCollection 2020. PLoS One. 2020. PMID: 31945099 Free PMC article.
-
The Genetic Basis of Pheromone Evolution in Moths.Annu Rev Entomol. 2016;61:99-117. doi: 10.1146/annurev-ento-010715-023638. Epub 2015 Nov 4. Annu Rev Entomol. 2016. PMID: 26565898 Review.
-
Regulation of pheromone biosynthesis in moths.Curr Opin Insect Sci. 2017 Dec;24:29-35. doi: 10.1016/j.cois.2017.09.002. Epub 2017 Sep 14. Curr Opin Insect Sci. 2017. PMID: 29208220 Review.
Cited by
-
Alcohol dehydrogenase 5 of Helicoverpa armigera interacts with the CYP6B6 promoter in response to 2-tridecanone.Insect Sci. 2020 Oct;27(5):1053-1066. doi: 10.1111/1744-7917.12720. Epub 2019 Sep 12. Insect Sci. 2020. PMID: 31454147 Free PMC article.
-
Analysis of natural female post-mating responses of Anopheles gambiae and Anopheles coluzzii unravels similarities and differences in their reproductive ecology.Sci Rep. 2018 Apr 26;8(1):6594. doi: 10.1038/s41598-018-24923-w. Sci Rep. 2018. PMID: 29700344 Free PMC article.
-
Transcriptome analysis identifies candidate genes in the biosynthetic pathway of sex pheromones from a zygaenid moth, Achelura yunnanensis (Lepidoptera: Zygaenidae).PeerJ. 2021 Dec 14;9:e12641. doi: 10.7717/peerj.12641. eCollection 2021. PeerJ. 2021. PMID: 34993022 Free PMC article.
-
Gypsy moth genome provides insights into flight capability and virus-host interactions.Proc Natl Acad Sci U S A. 2019 Jan 29;116(5):1669-1678. doi: 10.1073/pnas.1818283116. Epub 2019 Jan 14. Proc Natl Acad Sci U S A. 2019. PMID: 30642971 Free PMC article.
-
Comparative transcriptomics of the pheromone glands provides new insights into the differentiation of sex pheromone between two host populations of Chilo suppressalis.Sci Rep. 2020 Feb 26;10(1):3499. doi: 10.1038/s41598-020-60529-x. Sci Rep. 2020. PMID: 32103103 Free PMC article.
References
-
- Jurenka R: Insect pheromone biosynthesis. Top Curr Chem. 2004;239:97-132. doi:10.1007/b95450. - PubMed
-
- Zhu J-W, Millar J, Löfstedt C. Hormonal regulation of sex pheromone biosynthesis in the turnip moth, Agrotis segetum. Arch Insect Biochem Physiol. 1995;30(1):41–59. doi:10.1002/arch.940300104.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

