Tag mechanism as a strategy for the RNA replicase to resist parasites in the RNA world
- PMID: 28253281
- PMCID: PMC5333815
- DOI: 10.1371/journal.pone.0172702
Tag mechanism as a strategy for the RNA replicase to resist parasites in the RNA world
Abstract
The idea that life may have started with an "RNA world" is attractive. Wherein, a crucial event (perhaps at the very beginning of the scenario) should have been the emergence of a ribozyme that catalyzes its own replication, i.e., an RNA replicase. Although now there is experimental evidence supporting the chemical feasibility of such a ribozyme, the evolutionary dynamics of how the replicase could overcome the "parasite" problem (because other RNAs may also exploit this ribozyme) and thrive, as described in the scenario, remains unclear. It has been suggested that spatial limitation may have been important for the replicase to confront parasites. However, more studies showed that such a mechanism is not sufficient when this ribozyme's altruistic trait is taken into full consideration. "Tag mechanism", which means labeling the replicase with a short subsequence for recognition in replication, may be a further mechanism supporting the thriving of the replicase. However, because parasites may also "equip" themselves with the tag, it is far from clear whether the tag mechanism could take effect. Here, we conducted a computer simulation using a Monte-Carlo model to study the evolutionary dynamics surrounding the development of a tag-driven (polymerase-type) RNA replicase in the RNA world. We concluded that (1) with the tag mechanism the replicase could resist the parasites and become prosperous, (2) the main underlying reason should be that the parasitic molecules, especially those strong parasites, are more difficult to appear in the tag-driven system, and (3) the tag mechanism has a synergic effect with the spatial limitation mechanism-while the former provides "time" for the replicase to escape from parasites, the latter provides "space" for the replicase to escape. Notably, tags may readily serve as "control handles", and once the tag mechanism was exploited, the evolution towards complex life may have been much easier.
Conflict of interest statement
Figures



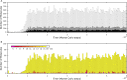

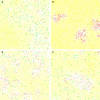

Similar articles
-
From molecular to cellular form: modeling the first major transition during the arising of life.BMC Evol Biol. 2019 Apr 3;19(1):84. doi: 10.1186/s12862-019-1412-5. BMC Evol Biol. 2019. PMID: 30943915 Free PMC article.
-
A simple template-dependent ligase ribozyme as the RNA replicase emerging first in the RNA world.Astrobiology. 2010 May;10(4):437-47. doi: 10.1089/ast.2009.0385. Astrobiology. 2010. PMID: 20528198
-
Nucleotide synthetase ribozymes may have emerged first in the RNA world.RNA. 2007 Nov;13(11):2012-9. doi: 10.1261/rna.658507. Epub 2007 Sep 18. RNA. 2007. PMID: 17878321 Free PMC article.
-
Intramolecular RNA replicase: possibly the first self-replicating molecule in the RNA world.Orig Life Evol Biosph. 2006 Aug;36(4):413-20. doi: 10.1007/s11084-005-9006-1. Epub 2006 Aug 15. Orig Life Evol Biosph. 2006. PMID: 16909330 Review.
-
Metabolically Coupled Replicator Systems: Overview of an RNA-world model concept of prebiotic evolution on mineral surfaces.J Theor Biol. 2015 Sep 21;381:39-54. doi: 10.1016/j.jtbi.2015.06.002. Epub 2015 Jun 15. J Theor Biol. 2015. PMID: 26087284 Review.
Cited by
-
What Does "the RNA World" Mean to "the Origin of Life"?Life (Basel). 2017 Nov 29;7(4):49. doi: 10.3390/life7040049. Life (Basel). 2017. PMID: 29186049 Free PMC article.
-
RNA World with Inhibitors.Entropy (Basel). 2024 Nov 23;26(12):1012. doi: 10.3390/e26121012. Entropy (Basel). 2024. PMID: 39766641 Free PMC article.
-
From molecular to cellular form: modeling the first major transition during the arising of life.BMC Evol Biol. 2019 Apr 3;19(1):84. doi: 10.1186/s12862-019-1412-5. BMC Evol Biol. 2019. PMID: 30943915 Free PMC article.
-
Multi-agent approach to sequence structure simulation in the RNA World hypothesis.PLoS One. 2020 Aug 28;15(8):e0238253. doi: 10.1371/journal.pone.0238253. eCollection 2020. PLoS One. 2020. PMID: 32857812 Free PMC article.
References
-
- Gilbert W. The RNA world. Nature. 1986; 319:618.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

