Ecological and Evolutionary Insights into Xanthomonas citri Pathovar Diversity
- PMID: 28258140
- PMCID: PMC5394309
- DOI: 10.1128/AEM.02993-16
Ecological and Evolutionary Insights into Xanthomonas citri Pathovar Diversity
Abstract
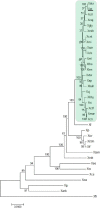
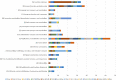
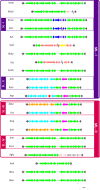
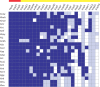
Citrus canker, caused by Xanthomonas citri pv. citri, is a serious disease of citrus plants worldwide. Earlier phylogenetic studies using housekeeping genes revealed that X. citri pv. citri is related to many other pathovars, which can be collectively referred as Xanthomonas citri pathovars (XCPs). From the present study, we report the genome sequences of 18 XCPs and compared them with four XCPs available in the public domain. In a tree based on phylogenomic marker genes, all the XCPs form a monophyletic cluster, suggesting their origin from a common ancestor. Phylogenomic analysis using the type strain further established that all the XCPs belong to one species. Clonal analysis of the core genome revealed the presence of two major lineages within this monophyletic cluster consisting of some clonal variants. Incidentally, the majority of these XCPs were first noticed in India, corroborating their clonal relationship and their common origin. Comparative analysis revealed an open pan-genome and the role of interstrain genomic flux of these XCPs since their diversification from a common ancestor. Even though there are wide variations in type III gene effectomes, we identified three core effectors which can be valuable in resistance-breeding programs. Overall, genomic examination of ecological relatives allowed us to dissect the tremendous genomic potential of X. citri species to rapidly evolve into specialized strains infecting diverse crop plants.IMPORTANCE Host specialization is one of the characteristic features of highly evolved pathogens such as the Xanthomonas group of phytopathogenic bacteria. Since the hosts involve staple crops and economically important fruits such as citrus, detailed understanding of the diversity and evolution of such strains infecting diverse plants is important for quarantine purposes. In the present study, we carried out genomic investigation of members of a phylogenetically and ecologically defined group of Xanthomonas strains pathogenic to diverse plants, including citrus. This group includes the oldest Xanthomonas pathovars and also recently emerged pathovars in a particular country where they are endemic. Our high-throughput genomic study has provided novel insights into the evolution of a unique lineage consisting of serious pathogens and their ecological relatives, suggesting the nature, scope, and pattern of rapid and recent diversification. Further, from the level of species to that of clonal variants, the study revealed interesting genomic patterns in diversification of a Xanthomonas lineage and perhaps will inspire careful study of the host range of the included pathovars.
Keywords: India; Xanthomonas; ecology; evolution; hypervariation; pathovar.
Copyright © 2017 American Society for Microbiology.
Figures


References
-
- Leyns F, De Cleene M, Swings J-G, De Ley J. 1984. The host range of the genus Xanthomonas. Botanical Rev 50:308–356. doi: 10.1007/BF02862635. - DOI
-
- Hayward A. 1993. The hosts of Xanthomonas, p 1–119. In Xanthomonas. Springer, New York, NY.
-
- Dye D, Bradbury J, Goto M, Hayward A, Lelliott R, Schroth M. 1980. International standards for naming pathovars of phytopathogenic bacteria and a list of pathovar names and pathotype strains. Rev Plant Pathol 59:153–168.
-
- Schaad NW, Postnikova E, Lacy GH, Sechler A, Agarkova I, Stromberg PE, Stromberg VK, Vidaver AK. 2005. Reclassification of Xanthomonas campestris pv. citri (ex Hasse 1915) Dye 1978 forms A, B/C/D, and E as X. smithii subsp. citri (ex Hasse) sp. nov. nom. rev. comb. nov., X. fuscans subsp. aurantifolii (ex Gabriel 1989) sp. nov. nom. rev. comb. nov., and X. alfalfae subsp. citrumelo (ex Riker and Jones) Gabriel et al., 1989 sp. nov. nom. rev. comb. nov.; X. campestris pv. malvacearum (ex Smith 1901) Dye 1978 as X. smithii subsp. smithii nov. comb. nov. nom. nov.; X. campestris pv. alfalfae (ex Riker and and Jones, 1935) Dye 1978 as X. alfalfae subsp. alfalfae (ex Riker et al., 1935) sp. nov. nom. rev.; and “var. fuscans” of X. campestris pv. phaseoli (ex Smith, 1987) Dye 1978 as X. fuscans subsp. fuscans sp. nov. Syst Appl Microbiol 28:494–518. doi: 10.1016/j.syapm.2005.03.017. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

