In silico identification of miRNAs and their target genes and analysis of gene co-expression network in saffron (Crocus sativus L. ) stigma
- PMID: 28261627
- PMCID: PMC5326487
In silico identification of miRNAs and their target genes and analysis of gene co-expression network in saffron (Crocus sativus L. ) stigma
Abstract
As an aromatic and colorful plant of substantive taste, saffron (Crocus sativus L.) owes such properties of matter to growing class of the secondary metabolites derived from the carotenoids, apocarotenoids. Regarding the critical role of microRNAs in secondary metabolic synthesis and the limited number of identified miRNAs in C. sativus, on the other hand, one may see the point how the characterization of miRNAs along with the corresponding target genes in C. sativus might expand our perspectives on the roles of miRNAs in carotenoid/apocarotenoid biosynthetic pathway. A computational analysis was used to identify miRNAs and their targets using EST (Expressed Sequence Tag) library from mature saffron stigmas. Then, a gene co- expression network was constructed to identify genes which are potentially involved in carotenoid/apocarotenoid biosynthetic pathways. EST analysis led to the identification of two putative miRNAs (miR414 and miR837-5p) along with the corresponding stem- looped precursors. To our knowledge, this is the first report on miR414 and miR837-5p in C. sativus. Co-expression network analysis indicated that miR414 and miR837-5p may play roles in C. sativus metabolic pathways and led to identification of candidate genes including six transcription factors and one protein kinase probably involved in carotenoid/apocarotenoid biosynthetic pathway. Presence of transcription factors, miRNAs and protein kinase in the network indicated multiple layers of regulation in saffron stigma. The candidate genes from this study may help unraveling regulatory networks underlying the carotenoid/apocarotenoid biosynthesis in saffron and designing metabolic engineering for enhanced secondary metabolites.
Keywords: Co-expression network; Crocus sativus; EST sequences analysis.
Figures
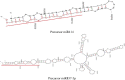
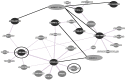
References
-
- Frusciante S, Diretto G, Bruno M, Ferrante P, Pietrella M, Prado-Cabrero A, Robio- Moraga A, Beyer P, Gomez-Gomez L, Al-Babili S, Giuliano G. Novel carotenoid cleavage dioxygenase catalyzes the first dedicated step in saffron crocin biosynthesis. Proc Natl Acad Sci USA. 2014;111:12246–12251. - PMC - PubMed
-
- Abdullaev FI, Espinosa-Aguirre JJ. Biomedical properties of saffron and its potential use in cancer therapy and chemoprevention trials. Cancer Detect Prev. 2004;28:426–432. - PubMed
-
- Gómez-Gómez L, Rubio-Moraga A, Ahrazem O. Understanding carotenoid metabolism in saffron stigmas: unraveling aroma and colour formation. Funct Plant Sci Biotechnol. 2010;4:56–63.
LinkOut - more resources
Full Text Sources
Research Materials
