Prevention of Bacterial Contamination of a Silica Matrix Containing Entrapped β-Galactosidase through the Action of Covalently Bound Lysozymes
- PMID: 28264511
- PMCID: PMC6155228
- DOI: 10.3390/molecules22030377
Prevention of Bacterial Contamination of a Silica Matrix Containing Entrapped β-Galactosidase through the Action of Covalently Bound Lysozymes
Abstract
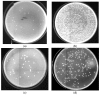
β-galactosidase was successfully encapsulated within an amino-functionalised silica matrix using a "fish-in-net" approach and molecular imprinting technique followed by covalent binding of lysozyme via a glutaraldehyde-based method. Transmission electron microscopy (TEM), X-ray diffraction (XRD), scanning electron microscopy (SEM), and Fourier transform infrared (FTIR) spectroscopy were used to characterise the silica matrix hosting the two enzymes. Both encapsulated β-galactosidase and bound lysozyme exhibited high enzymatic activities and outstanding operational stability in model reactions. Moreover, enzyme activities of the co-immobilised enzymes did not obviously change relative to enzymes immobilised separately. In antibacterial tests, bound lysozyme exhibited 95.5% and 89.6% growth inhibition of Staphylococcus aureus ATCC (American type culture collection) 653 and Escherichia coli ATCC 1122, respectively. In milk treated with co-immobilised enzymes, favourable results were obtained regarding reduction of cell viability and high lactose hydrolysis rate. In addition, when both co-immobilised enzymes were employed to treat milk, high operational and storage stabilities were observed. The results demonstrate that the use of co-immobilised enzymes holds promise as an industrial strategy for producing low lactose milk to benefit people with lactose intolerance.
Keywords: covalent binding; encapsulation; lysozyme; β-galactosidase.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Benavente R., Pessela B., Curiel J., de las Rivas B., Muñoz R., Guisán J., Mancheño J., Cardelle-Cobas A., Ruiz-Matute A., Corzo N. Improving properties of a novel β-galactosidase from Lactobacillus plantarum by covalent immobilization. Molecules. 2015;20:7874. doi: 10.3390/molecules20057874. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

