Systematic substitutions at BLIP position 50 result in changes in binding specificity for class A β-lactamases
- PMID: 28264645
- PMCID: PMC5340008
- DOI: 10.1186/s12858-017-0077-1
Systematic substitutions at BLIP position 50 result in changes in binding specificity for class A β-lactamases
Abstract
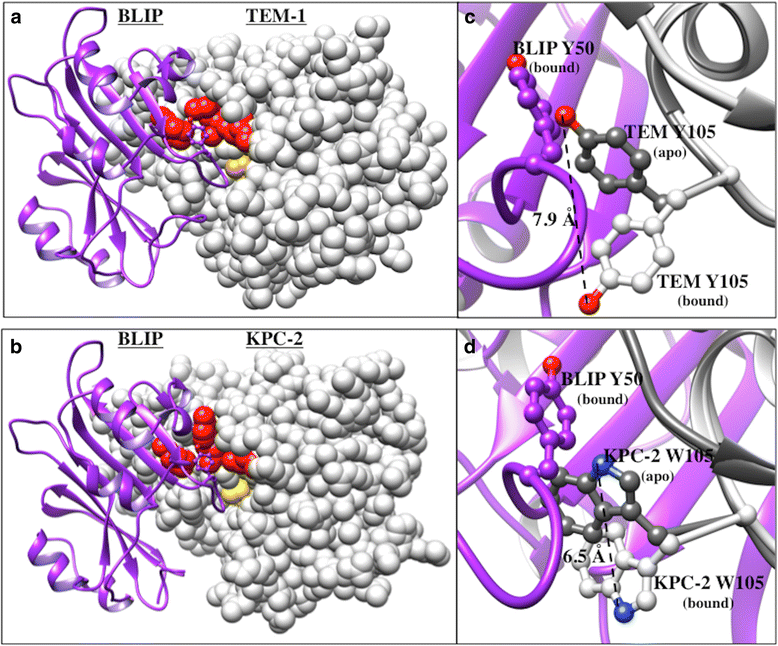
Background: The production of β-lactamases by bacteria is the most common mechanism of resistance to the widely prescribed β-lactam antibiotics. β-lactamase inhibitory protein (BLIP) competitively inhibits class A β-lactamases via two binding loops that occlude the active site. It has been shown that BLIP Tyr50 is a specificity determinant in that substitutions at this position result in large differential changes in the relative affinity of BLIP for class A β-lactamases.
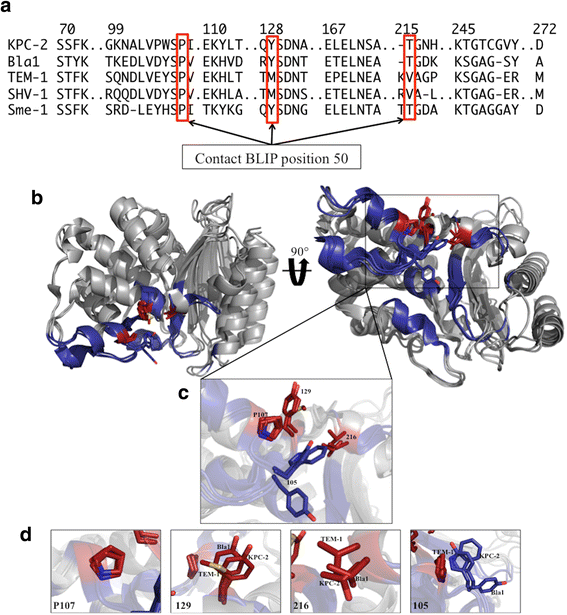
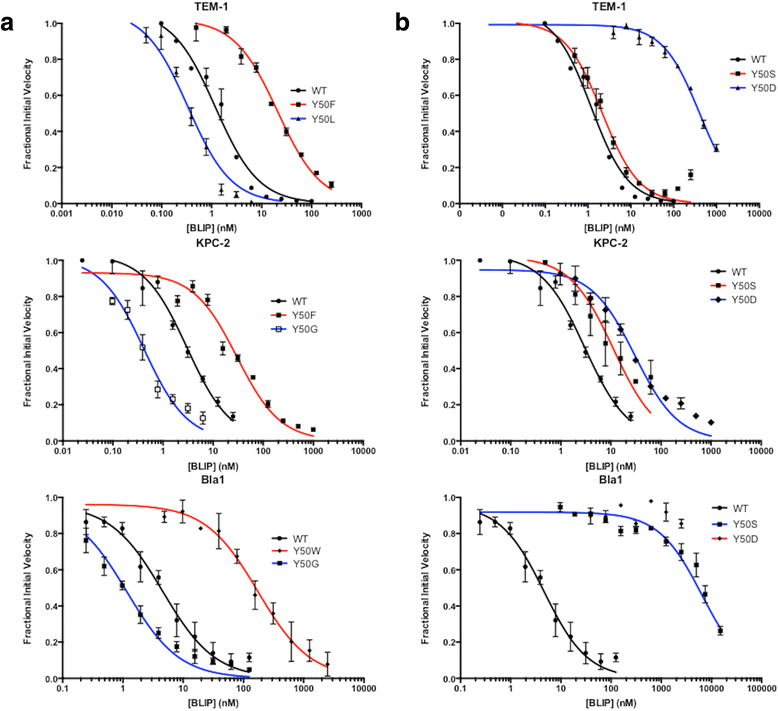
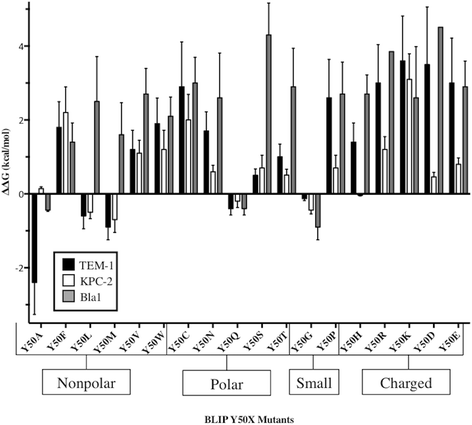
Results: In this study, the effect of systematic substitutions at BLIP position 50 on binding to class A β-lactamases was examined to further explore the role of BLIP Tyr50 in modulating specificity. The results indicate the sequence requirements at position 50 are widely different depending on the target β-lactamase. Stringent sequence requirements were observed at Tyr50 for binding Bacillus anthracis Bla1 while moderate requirements for binding TEM-1 and relaxed requirements for binding KPC-2 β-lactamase were seen. These findings cannot be easily rationalized based on the β-lactamase residues in direct contact with BLIP Tyr50 since they are identical for Bla1 and KPC-2 suggesting that differences in the BLIP-β-lactamase interface outside the local environment of Tyr50 influence the effect of substitutions.
Conclusions: Results from this study and previous studies suggest that substitutions at BLIP Tyr50 may induce changes at the interface outside its local environment and point to the complexity of predicting the impact of substitutions at a protein-protein interaction interface.
Keywords: Beatmusic; Binding specificity; Systematic substitutions; β-lactamase.
Figures
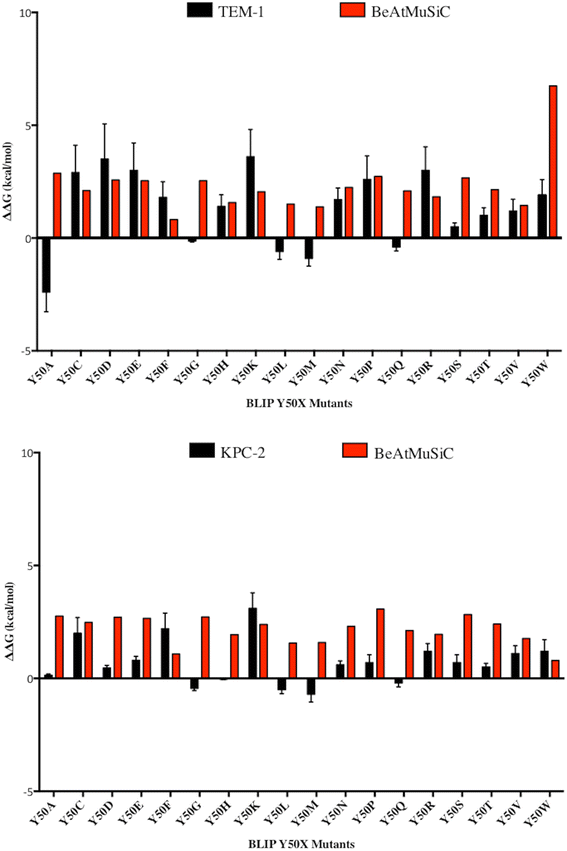
References
-
- Kuzu G, Keskin O, Gursoy A, Nussinov R. Constructing structural networks of signaling pathways on the proteome scale. Curr Opin Struc Biol. sciencedirect; 2012;22:367–377 - PubMed
-
- Jubb H, Higueruelo A, Winter A, Blundell T. Structural biology and drug discovery for protein–protein interactions. Trends Pharmacol Sci. 2012;33:241–248. - PubMed
-
- Selzer T, Albeck S, Schreiber G. Rational design of faster associating and tighter binding protein complexes. Nat. Struct. Biol. 2000;7:537–41. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

