LoTo: a graphlet based method for the comparison of local topology between gene regulatory networks
- PMID: 28265516
- PMCID: PMC5333545
- DOI: 10.7717/peerj.3052
LoTo: a graphlet based method for the comparison of local topology between gene regulatory networks
Abstract
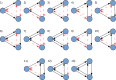
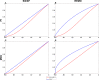
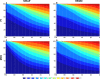
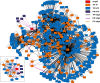
One of the main challenges of the post-genomic era is the understanding of how gene expression is controlled. Changes in gene expression lay behind diverse biological phenomena such as development, disease and the adaptation to different environmental conditions. Despite the availability of well-established methods to identify these changes, tools to discern how gene regulation is orchestrated are still required. The regulation of gene expression is usually depicted as a Gene Regulatory Network (GRN) where changes in the network structure (i.e., network topology) represent adjustments of gene regulation. Like other networks, GRNs are composed of basic building blocks; small induced subgraphs called graphlets. Here we present LoTo, a novel method that using Graphlet Based Metrics (GBMs) identifies topological variations between different states of a GRN. Under our approach, different states of a GRN are analyzed to determine the types of graphlet formed by all triplets of nodes in the network. Subsequently, graphlets occurring in a state of the network are compared to those formed by the same three nodes in another version of the network. Once the comparisons are performed, LoTo applies metrics from binary classification problems calculated on the existence and absence of graphlets to assess the topological similarity between both network states. Experiments performed on randomized networks demonstrate that GBMs are more sensitive to topological variation than the same metrics calculated on single edges. Additional comparisons with other common metrics demonstrate that our GBMs are capable to identify nodes whose local topology changes between different states of the network. Notably, due to the explicit use of graphlets, LoTo captures topological variations that are disregarded by other approaches. LoTo is freely available as an online web server at http://dlab.cl/loto.
Keywords: Differential analysis; Gene Regulatory Network; Graphlet; Metric.
Conflict of interest statement
Tomas Perez-Acle is an Academic Editor for PeerJ.
Figures
Similar articles
-
Graphlet Based Metrics for the Comparison of Gene Regulatory Networks.PLoS One. 2016 Oct 3;11(10):e0163497. doi: 10.1371/journal.pone.0163497. eCollection 2016. PLoS One. 2016. PMID: 27695050 Free PMC article.
-
IncGraph: Incremental graphlet counting for topology optimisation.PLoS One. 2018 Apr 26;13(4):e0195997. doi: 10.1371/journal.pone.0195997. eCollection 2018. PLoS One. 2018. PMID: 29698494 Free PMC article.
-
Graphlet Laplacians for topology-function and topology-disease relationships.Bioinformatics. 2019 Dec 15;35(24):5226-5234. doi: 10.1093/bioinformatics/btz455. Bioinformatics. 2019. PMID: 31192358
-
Exploiting graphlet decomposition to explain the structure of complex networks: the GHuST framework.Sci Rep. 2020 Jul 30;10(1):12884. doi: 10.1038/s41598-020-69795-1. Sci Rep. 2020. PMID: 32732972 Free PMC article.
-
[Gene regulatory network of hepatocellular carcinoma: a review].Sheng Wu Gong Cheng Xue Bao. 2016 Oct 25;32(10):1322-1331. doi: 10.13345/j.cjb.160045. Sheng Wu Gong Cheng Xue Bao. 2016. PMID: 29027443 Review. Chinese.
Cited by
-
Automated generation of context-specific gene regulatory networks with a weighted approach in Drosophila melanogaster.Interface Focus. 2021 Jun 11;11(4):20200076. doi: 10.1098/rsfs.2020.0076. eCollection 2021 Jun. Interface Focus. 2021. PMID: 34123358 Free PMC article.
-
Network subgraph-based approach for analyzing and comparing molecular networks.PeerJ. 2022 May 3;10:e13137. doi: 10.7717/peerj.13137. eCollection 2022. PeerJ. 2022. PMID: 35529499 Free PMC article.
-
Homology-based reconstruction of regulatory networks for bacterial and archaeal genomes.Front Microbiol. 2022 Jul 19;13:923105. doi: 10.3389/fmicb.2022.923105. eCollection 2022. Front Microbiol. 2022. PMID: 35928164 Free PMC article.
-
Network-Based Approaches Reveal Potential Therapeutic Targets for Host-Directed Antileishmanial Therapy Driving Drug Repurposing.Microbiol Spectr. 2021 Oct 31;9(2):e0101821. doi: 10.1128/Spectrum.01018-21. Epub 2021 Oct 20. Microbiol Spectr. 2021. PMID: 34668739 Free PMC article.
-
Dissecting molecular network structures using a network subgraph approach.PeerJ. 2020 Aug 6;8:e9556. doi: 10.7717/peerj.9556. eCollection 2020. PeerJ. 2020. PMID: 33005483 Free PMC article.
References
-
- Aparício DO, Ribeiro PMP, Silva FMA. Network comparison using directed graphlets. 20151511.01964
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

