Efficient and rapid generation of large genomic variants in rats and mice using CRISMERE
- PMID: 28266534
- PMCID: PMC5339700
- DOI: 10.1038/srep43331
Efficient and rapid generation of large genomic variants in rats and mice using CRISMERE
Abstract
Modelling Down syndrome (DS) in mouse has been crucial for the understanding of the disease and the evaluation of therapeutic targets. Nevertheless, the modelling so far has been limited to the mouse and, even in this model, generating duplication of genomic regions has been labour intensive and time consuming. We developed the CRISpr MEdiated REarrangement (CRISMERE) strategy, which takes advantage of the CRISPR/Cas9 system, to generate most of the desired rearrangements from a single experiment at much lower expenses and in less than 9 months. Deletions, duplications, and inversions of genomic regions as large as 24.4 Mb in rat and mouse founders were observed and germ line transmission was confirmed for fragment as large as 3.6 Mb. Interestingly we have been able to recover duplicated regions from founders in which we only detected deletions. CRISMERE is even more powerful than anticipated it allows the scientific community to manipulate the rodent and probably other genomes in a fast and efficient manner which was not possible before.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




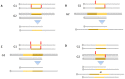
References
-
- Dierssen M., Herault Y. & Estivill X. Aneuploidy: from a physiological mechanism of variance to Down syndrome. Physiol Rev 89, 887–920 (2009). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

