Misrouting of v-ATPase subunit V0a1 dysregulates lysosomal acidification in a neurodegenerative lysosomal storage disease model
- PMID: 28266544
- PMCID: PMC5344305
- DOI: 10.1038/ncomms14612
Misrouting of v-ATPase subunit V0a1 dysregulates lysosomal acidification in a neurodegenerative lysosomal storage disease model
Abstract
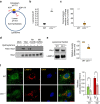
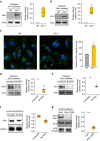
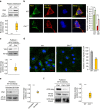
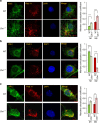
Defective lysosomal acidification contributes to virtually all lysosomal storage disorders (LSDs) and to common neurodegenerative diseases like Alzheimer's and Parkinson's. Despite its fundamental importance, the mechanism(s) underlying this defect remains unclear. The v-ATPase, a multisubunit protein complex composed of cytosolic V1-sector and lysosomal membrane-anchored V0-sector, regulates lysosomal acidification. Mutations in the CLN1 gene, encoding PPT1, cause a devastating neurodegenerative LSD, INCL. Here we report that in Cln1-/- mice, which mimic INCL, reduced v-ATPase activity correlates with elevated lysosomal pH. Moreover, v-ATPase subunit a1 of the V0 sector (V0a1) requires palmitoylation for interacting with adaptor protein-2 (AP-2) and AP-3, respectively, for trafficking to the lysosomal membrane. Notably, treatment of Cln1-/- mice with a thioesterase (Ppt1)-mimetic, NtBuHA, ameliorated this defect. Our findings reveal an unanticipated role of Cln1 in regulating lysosomal targeting of V0a1 and suggest that varying factors adversely affecting v-ATPase function dysregulate lysosomal acidification in other LSDs and common neurodegenerative diseases.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



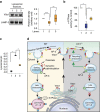
References
-
- de Duve C. & Wattiaux R. Functions of lysosomes. Annu. Rev. Physiol. 28, 435–492 (1966). - PubMed
-
- Mellman I., Fuchs R. & Helenius A. Acidification of the endocytic and exocytic pathways. Annu. Rev. Biochem. 55, 663–700 (1986). - PubMed
-
- Futerman A. H. & van Meer G. The cell biology of lysosomal storage disorders. Nat. Rev. Mol. Cell Biol. 5, 554–565 (2004). - PubMed
-
- Holopainen J. M., Saarikoski J., Kinnunen P. K. & Järvelä I. Elevated lysosomal pH in neuronal ceroid lipofuscinoses. Eur. J. Biochem. 268, 5851–5856 (2001). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

