A self-perpetuating repressive state of a viral replication protein blocks superinfection by the same virus
- PMID: 28267773
- PMCID: PMC5357057
- DOI: 10.1371/journal.ppat.1006253
A self-perpetuating repressive state of a viral replication protein blocks superinfection by the same virus
Abstract
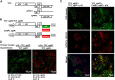
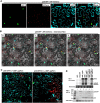
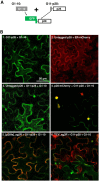
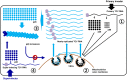
Diverse animal and plant viruses block the re-infection of host cells by the same or highly similar viruses through superinfection exclusion (SIE), a widely observed, yet poorly understood phenomenon. Here we demonstrate that SIE of turnip crinkle virus (TCV) is exclusively determined by p28, one of the two replication proteins encoded by this virus. p28 expressed from a TCV replicon exerts strong SIE to a different TCV replicon. Transiently expressed p28, delivered simultaneously with, or ahead of, a TCV replicon, largely recapitulates this repressive activity. Interestingly, p28-mediated SIE is dramatically enhanced by C-terminally fused epitope tags or fluorescent proteins, but weakened by N-terminal modifications, and it inversely correlates with the ability of p28 to complement the replication of a p28-defective TCV replicon. Strikingly, p28 in SIE-positive cells forms large, mobile punctate inclusions that trans-aggregate a non-coalescing, SIE-defective, yet replication-competent p28 mutant. These results support a model postulating that TCV SIE is caused by the formation of multimeric p28 complexes capable of intercepting fresh p28 monomers translated from superinfector genomes, thereby abolishing superinfector replication. This model could prove to be applicable to other RNA viruses, and offer novel targets for antiviral therapy.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
Superinfection Exclusion by p28 of Turnip Crinkle Virus Is Separable from Its Replication Function.Mol Plant Microbe Interact. 2020 Feb;33(2):364-375. doi: 10.1094/MPMI-09-19-0258-R. Epub 2019 Dec 27. Mol Plant Microbe Interact. 2020. PMID: 31880982
-
Repression of turnip crinkle virus replication by its replication protein p88.Virology. 2019 Jan 2;526:165-172. doi: 10.1016/j.virol.2018.10.024. Epub 2018 Nov 2. Virology. 2019. PMID: 30391806
-
Replication-Dependent Biogenesis of Turnip Crinkle Virus Long Noncoding RNAs.J Virol. 2021 Aug 25;95(18):e0016921. doi: 10.1128/JVI.00169-21. Epub 2021 Aug 25. J Virol. 2021. PMID: 34160262 Free PMC article.
-
A New Mechanistic Model for Viral Cross Protection and Superinfection Exclusion.Front Plant Sci. 2018 Jan 25;9:40. doi: 10.3389/fpls.2018.00040. eCollection 2018. Front Plant Sci. 2018. PMID: 29422912 Free PMC article. Review.
-
3'UTRs of carmoviruses.Virus Res. 2015 Aug 3;206:27-36. doi: 10.1016/j.virusres.2015.01.023. Epub 2015 Feb 4. Virus Res. 2015. PMID: 25662021 Review.
Cited by
-
Developing reverse genetics systems of northern cereal mosaic virus to reveal superinfection exclusion of two cytorhabdoviruses in barley plants.Mol Plant Pathol. 2022 May;23(5):749-756. doi: 10.1111/mpp.13188. Epub 2022 Feb 6. Mol Plant Pathol. 2022. PMID: 35124878 Free PMC article.
-
Stable and Broad Spectrum Cross-Protection Against Pepino Mosaic Virus Attained by Mixed Infection.Front Plant Sci. 2018 Dec 6;9:1810. doi: 10.3389/fpls.2018.01810. eCollection 2018. Front Plant Sci. 2018. PMID: 30574159 Free PMC article.
-
Bottleneck, Isolate, Amplify, Select (BIAS) as a mechanistic framework for intracellular population dynamics of positive-sense RNA viruses.Virus Evol. 2020 Nov 12;6(2):veaa086. doi: 10.1093/ve/veaa086. eCollection 2020 Jul. Virus Evol. 2020. PMID: 33343926 Free PMC article. Review.
-
The fitness consequences of coinfection and reassortment for segmented viruses depend upon viral genetic structure.bioRxiv [Preprint]. 2025 Jul 26:2025.07.22.666171. doi: 10.1101/2025.07.22.666171. bioRxiv. 2025. PMID: 40777271 Free PMC article. Preprint.
-
P3 and NIa-Pro of Turnip Mosaic Virus Are Independent Elicitors of Superinfection Exclusion.Viruses. 2023 Jun 28;15(7):1459. doi: 10.3390/v15071459. Viruses. 2023. PMID: 37515147 Free PMC article.
References
-
- Simon KO, Cardamone JJ Jr, Whitaker-Dowling A, Youngner JS, Widnell CC (1990) Cellular mechanisms in the superinfection exclusion of vesicular stomatitis virus. Virology 177:375–379. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

