Adaptation of A-to-I RNA editing in Drosophila
- PMID: 28282384
- PMCID: PMC5365144
- DOI: 10.1371/journal.pgen.1006648
Adaptation of A-to-I RNA editing in Drosophila
Abstract
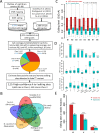
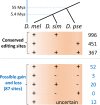
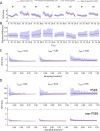
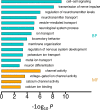
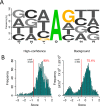
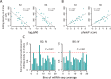
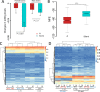
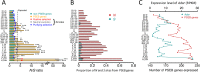
Adenosine-to-inosine (A-to-I) editing is hypothesized to facilitate adaptive evolution by expanding proteomic diversity through an epigenetic approach. However, it is challenging to provide evidences to support this hypothesis at the whole editome level. In this study, we systematically characterized 2,114 A-to-I RNA editing sites in female and male brains of D. melanogaster, and nearly half of these sites had events evolutionarily conserved across Drosophila species. We detected strong signatures of positive selection on the nonsynonymous editing sites in Drosophila brains, and the beneficial editing sites were significantly enriched in genes related to chemical and electrical neurotransmission. The signal of adaptation was even more pronounced for the editing sites located in X chromosome or for those commonly observed across Drosophila species. We identified a set of gene candidates (termed "PSEB" genes) that had nonsynonymous editing events favored by natural selection. We presented evidence that editing preferentially increased mutation sequence space of evolutionarily conserved genes, which supported the adaptive evolution hypothesis of editing. We found prevalent nonsynonymous editing sites that were favored by natural selection in female and male adults from five strains of D. melanogaster. We showed that temperature played a more important role than gender effect in shaping the editing levels, although the effect of temperature is relatively weaker compared to that of species effect. We also explored the relevant factors that shape the selective patterns of the global editomes. Altogether we demonstrated that abundant nonsynonymous editing sites in Drosophila brains were adaptive and maintained by natural selection during evolution. Our results shed new light on the evolutionary principles and functional consequences of RNA editing.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
Linkage of A-to-I RNA Editing in Metazoans and the Impact on Genome Evolution.Mol Biol Evol. 2018 Jan 1;35(1):132-148. doi: 10.1093/molbev/msx274. Mol Biol Evol. 2018. PMID: 29048557 Free PMC article.
-
New comparative genomic evidence supporting the proteomic diversification role of A-to-I RNA editing in insects.Mol Genet Genomics. 2024 Apr 20;299(1):46. doi: 10.1007/s00438-024-02141-6. Mol Genet Genomics. 2024. PMID: 38642133
-
Conserved A-to-I RNA editing with non-conserved recoding expands the candidates of functional editing sites.Fly (Austin). 2024 Dec;18(1):2367359. doi: 10.1080/19336934.2024.2367359. Epub 2024 Jun 18. Fly (Austin). 2024. PMID: 38889318 Free PMC article.
-
The Many Roles of A-to-I RNA Editing in Animals: Functional or Adaptive?Front Biosci (Landmark Ed). 2023 Oct 20;28(10):256. doi: 10.31083/j.fbl2810256. Front Biosci (Landmark Ed). 2023. PMID: 37919076 Review.
-
Evolutionary driving forces of A-to-I editing in metazoans.Wiley Interdiscip Rev RNA. 2022 Jan;13(1):e1666. doi: 10.1002/wrna.1666. Epub 2021 May 16. Wiley Interdiscip Rev RNA. 2022. PMID: 33998151 Review.
Cited by
-
Adaptive Proteome Diversification by Nonsynonymous A-to-I RNA Editing in Coleoid Cephalopods.Mol Biol Evol. 2021 Aug 23;38(9):3775-3788. doi: 10.1093/molbev/msab154. Mol Biol Evol. 2021. PMID: 34022057 Free PMC article.
-
The evolutionary arms race between transposable elements and piRNAs in Drosophila melanogaster.BMC Evol Biol. 2020 Jan 28;20(1):14. doi: 10.1186/s12862-020-1580-3. BMC Evol Biol. 2020. PMID: 31992188 Free PMC article.
-
Human A-to-I RNA editing SNP loci are enriched in GWAS signals for autoimmune diseases and under balancing selection.Genome Biol. 2020 Nov 30;21(1):288. doi: 10.1186/s13059-020-02205-x. Genome Biol. 2020. PMID: 33256812 Free PMC article.
-
Illuminating spatial A-to-I RNA editing signatures within the Drosophila brain.Proc Natl Acad Sci U S A. 2019 Feb 5;116(6):2318-2327. doi: 10.1073/pnas.1811768116. Epub 2019 Jan 18. Proc Natl Acad Sci U S A. 2019. PMID: 30659150 Free PMC article.
-
Linkage of A-to-I RNA Editing in Metazoans and the Impact on Genome Evolution.Mol Biol Evol. 2018 Jan 1;35(1):132-148. doi: 10.1093/molbev/msx274. Mol Biol Evol. 2018. PMID: 29048557 Free PMC article.
References
-
- Fisher RA. The genetical theory of natural selection. Oxford: Clarendon Press; 1930.
-
- Hedrick PW. Antagonistic pleiotropy and genetic polymorphism: a perspective. Heredity. 1999;82(2):126–33.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

