Nitrogen Metabolism Genes from Temperate Marine Sediments
- PMID: 28283802
- PMCID: PMC5405112
- DOI: 10.1007/s10126-017-9741-0
Nitrogen Metabolism Genes from Temperate Marine Sediments
Abstract
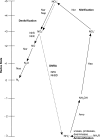
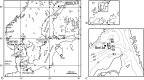
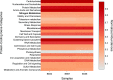
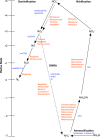
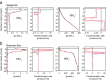
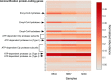
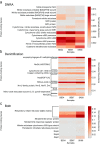
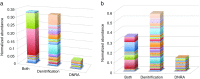
In this study, we analysed metagenomes along with biogeochemical profiles from Skagerrak (SK) and Bothnian Bay (BB) sediments, to trace the prevailing nitrogen pathways. NO3- was present in the top 5 cm below the sediment-water interface at both sites. NH4+ increased with depth below 5 cm where it overlapped with the NO3- zone. Steady-state modelling of NO3- and NH4+ porewater profiles indicates zones of net nitrogen species transformations. Bacterial protease and hydratase genes appeared to make up the bulk of total ammonification genes. Genes involved in ammonia oxidation (amo, hao), denitrification (nir, nor), dissimilatory NO3- reduction to NH4+ (nfr and otr) and in both of the latter two pathways (nar, nap) were also present. Results show ammonia-oxidizing bacteria (AOB) and ammonia-oxidizing archaea (AOA) are similarly abundant in both sediments. Also, denitrification genes appeared more abundant than DNRA genes. 16S rRNA gene analysis showed that the relative abundance of the nitrifying group Nitrosopumilales and other groups involved in nitrification and denitrification (Nitrobacter, Nitrosomonas, Nitrospira, Nitrosococcus and Nitrosomonas) appeared less abundant in SK sediments compared to BB sediments. Beggiatoa and Thiothrix 16S rRNA genes were also present, suggesting chemolithoautotrophic NO3- reduction to NO2- or NH4+ as a possible pathway. Our results show the metabolic potential for ammonification, nitrification, DNRA and denitrification activities in North Sea and Baltic Sea sediments.
Keywords: Marine; Metagenome; Nitrogen; Sediments.
Figures


References
-
- Algar CK, Vallino JJ. Predicting microbial nitrate reduction pathways in coastal sediments. Aquat Microb Ecol. 2014;71:223–238. doi: 10.3354/ame01678. - DOI
-
- Baggs E, Phillipot L (2011) Nitrous oxide production in the terrestrial environment. In: Moir JWB (ed) Nitrogen cycling in bacteria: molecular analysis. Caister Academic Press, Norfolk
-
- Bale NJ, Villanueva L, Hopmans EC, Schouten S, Sinninghe Damsté JS. Different seasonality of pelagic and benthic Thaumarchaeota in the North Sea. Biogeosciences. 2013;10:7195–7206. doi: 10.5194/bg-10-7195-2013. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
