Dynamic Role of trans Regulation of Gene Expression in Relation to Complex Traits
- PMID: 28285768
- PMCID: PMC5384035
- DOI: 10.1016/j.ajhg.2017.02.003
Dynamic Role of trans Regulation of Gene Expression in Relation to Complex Traits
Erratum in
-
Dynamic Role of trans Regulation of Gene Expression in Relation to Complex Traits.Am J Hum Genet. 2017 Jun 1;100(6):985-986. doi: 10.1016/j.ajhg.2017.05.002. Am J Hum Genet. 2017. PMID: 28575653 Free PMC article. No abstract available.
Abstract
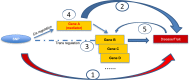
Identifying causal genetic variants and understanding their mechanisms of effect on traits remains a challenge in genome-wide association studies (GWASs). In particular, how genetic variants (i.e., trans-eQTLs) affect expression of remote genes (i.e., trans-eGenes) remains unknown. We hypothesized that some trans-eQTLs regulate expression of distant genes by altering the expression of nearby genes (cis-eGenes). Using published GWAS datasets with 39,165 single-nucleotide polymorphisms (SNPs) associated with 1,960 traits, we explored whole blood gene expression associations of trait-associated SNPs in 5,257 individuals from the Framingham Heart Study. We identified 2,350 trans-eQTLs (at p < 10-7); more than 80% of them were found to have cis-associated eGenes. Mediation testing suggested that for 35% of trans-eQTL-trans-eGene pairs in different chromosomes and 90% pairs in the same chromosome, the disease-associated SNP may alter expression of the trans-eGene via cis-eGene expression. In addition, we identified 13 trans-eQTL hotspots, affecting from ten to hundreds of genes, suggesting the existence of master genetic regulators. Using causal inference testing, we searched causal variants across eight cardiometabolic traits (BMI, systolic and diastolic blood pressure, LDL cholesterol, HDL cholesterol, total cholesterol, triglycerides, and fasting blood glucose) and identified several cis-eGenes (ALDH2 for systolic and diastolic blood pressure, MCM6 and DARS for total cholesterol, and TRIB1 for triglycerides) that were causal mediators for the corresponding traits, as well as examples of trans-mediators (TAGAP for LDL cholesterol). The finding of extensive evidence of genome-wide mediation effects suggests a critical role of cryptic gene regulation underlying many disease traits.
Keywords: GWAS; cardiometabolic traits; causal variants; eQTLs; hotspots; mediation; trans.
Published by Elsevier Inc.
Figures



Comment in
-
Disease genomics: Transitioning from association to causation with eQTLs.Nat Rev Genet. 2017 May;18(5):271. doi: 10.1038/nrg.2017.22. Epub 2017 Mar 27. Nat Rev Genet. 2017. PMID: 28344342 No abstract available.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

