Ordering the mob: Insights into replicon and MOB typing schemes from analysis of a curated dataset of publicly available plasmids
- PMID: 28286183
- PMCID: PMC5466382
- DOI: 10.1016/j.plasmid.2017.03.002
Ordering the mob: Insights into replicon and MOB typing schemes from analysis of a curated dataset of publicly available plasmids
Abstract
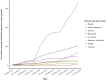
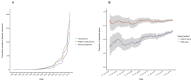
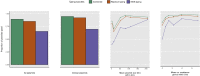
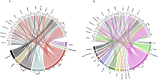
Plasmid typing can provide insights into the epidemiology and transmission of plasmid-mediated antibiotic resistance. The principal plasmid typing schemes are replicon typing and MOB typing, which utilize variation in replication loci and relaxase proteins respectively. Previous studies investigating the proportion of plasmids assigned a type by these schemes ('typeability') have yielded conflicting results; moreover, thousands of plasmid sequences have been added to NCBI in recent years, without consistent annotation to indicate which sequences represent complete plasmids. Here, a curated dataset of complete Enterobacteriaceae plasmids from NCBI was compiled, and used to assess the typeability and concordance of in silico replicon and MOB typing schemes. Concordance was assessed at hierarchical replicon type resolutions, from replicon family-level to plasmid multilocus sequence type (pMLST)-level, where available. We found that 85% and 65% of the curated plasmids could be replicon and MOB typed, respectively. Overall, plasmid size and the number of resistance genes were significant independent predictors of replicon and MOB typing success. We found some degree of non-concordance between replicon families and MOB types, which was only partly resolved when partitioning plasmids into finer-resolution groups (replicon and pMLST types). In some cases, non-concordance was attributed to ambiguous boundaries between MOBP and MOBQ types; in other cases, backbone mosaicism was considered a more plausible explanation. β-lactamase resistance genes tended not to show fidelity to a particular plasmid type, though some previously reported associations were supported. Overall, replicon and MOB typing schemes are likely to continue playing an important role in plasmid analysis, but their performance is constrained by the diverse and dynamic nature of plasmid genomes.
Keywords: Antibiotic resistance; MOB typing; Plasmid database; Plasmid multilocus sequence typing; Replicon typing.
Copyright © 2017 The Authors. Published by Elsevier Inc. All rights reserved.
Figures


References
-
- Akiba M., Sekizuka T., Yamashita A., Kuroda M., Fujii Y., Murata M. Vol. 60. 2016. Distribution and Relationships of Antimicrobial Resistance Determinants Among Extended-Spectrum-Cephalosporin-Resistant or Carbapenem-Resistant Escherichia coli Isolates From Rivers and Sewage Treatment Plants in India; pp. 2972–2980. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

