Crystal structures reveal N-terminal Domain of Arabidopsis thaliana ClpD to be highly divergent from that of ClpC1
- PMID: 28287170
- PMCID: PMC5347014
- DOI: 10.1038/srep44366
Crystal structures reveal N-terminal Domain of Arabidopsis thaliana ClpD to be highly divergent from that of ClpC1
Abstract
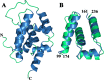
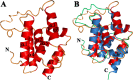
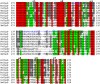
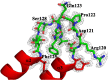
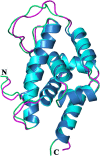
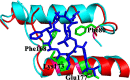
The caseinolytic protease machinery associated chaperone protein ClpC is known to be present in bacteria, plants and other eukaryotes, whereas ClpD is unique to plants. Plant ClpC and ClpD proteins get localized into chloroplast stroma. Herein, we report high resolution crystal structures of the N-terminal domain of Arabidopsis thaliana ClpC1 and ClpD. Surprisingly, AtClpD, but not AtClpC1, deviates from the typical N-terminal repeat domain organization of known Clp chaperones and have only seven α-helices, instead of eight. In addition, the loop connecting the two halves of AtClpD NTD is longer and covers the region which in case of AtClpC1 is thought to contribute to adaptor protein interaction. Taken together, the N-terminal domain of AtClpD has a divergent structural organization compared to any known Clp chaperones which hints towards its specific role during plant stress conditions, as opposed to that in the maintenance of chloroplastic homeostasis by AtClpC1. Conservation of residues in the NTD that are responsible for the binding of the cyclic peptide activator - Cyclomarin A, as reported for mycobacterial ClpC1 suggests that the peptide could be used as an activator to both AtClpC1 and AtClpD, which could be useful in their detailed in vitro functional characterization.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




Similar articles
-
Recombinant expression, purification and SAXS analysis of Arabidopsis thaliana ClpC1.Int J Biol Macromol. 2021 Jan 15;167:1273-1280. doi: 10.1016/j.ijbiomac.2020.11.081. Epub 2020 Nov 13. Int J Biol Macromol. 2021. PMID: 33189753
-
In Vivo Trapping of Proteins Interacting with the Chloroplast CLPC1 Chaperone: Potential Substrates and Adaptors.J Proteome Res. 2019 Jun 7;18(6):2585-2600. doi: 10.1021/acs.jproteome.9b00112. Epub 2019 May 22. J Proteome Res. 2019. PMID: 31070379
-
Chloroplastic Hsp100 chaperones ClpC2 and ClpD interact in vitro with a transit peptide only when it is located at the N-terminus of a protein.BMC Plant Biol. 2012 Apr 30;12:57. doi: 10.1186/1471-2229-12-57. BMC Plant Biol. 2012. PMID: 22545953 Free PMC article.
-
Uvr motifs regulate the chloroplast Clp chaperone-protease system.Trends Plant Sci. 2025 Mar;30(3):269-282. doi: 10.1016/j.tplants.2024.09.015. Epub 2024 Oct 24. Trends Plant Sci. 2025. PMID: 39448301 Review.
-
Arabidopsis thaliana J-class heat shock proteins: cellular stress sensors.Funct Integr Genomics. 2009 Nov;9(4):433-46. doi: 10.1007/s10142-009-0132-0. Epub 2009 Jul 25. Funct Integr Genomics. 2009. PMID: 19633874 Review.
Cited by
-
Structure, function, and substrates of Clp AAA+ protease systems in cyanobacteria, plastids, and apicoplasts: A comparative analysis.J Biol Chem. 2021 Jan-Jun;296:100338. doi: 10.1016/j.jbc.2021.100338. Epub 2021 Jan 23. J Biol Chem. 2021. PMID: 33497624 Free PMC article. Review.
-
Deciphering the mechanism and function of Hsp100 unfoldases from protein structure.Biochem Soc Trans. 2022 Dec 16;50(6):1725-1736. doi: 10.1042/BST20220590. Biochem Soc Trans. 2022. PMID: 36454589 Free PMC article.
References
-
- Adam Z. & Clarke A. K. Cutting edge of chloroplast proteolysis. Trends Plant Sci. 7, 451–456, doi: S1360-1385(02)02326-9 (2002). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

