Increase of Fungal Pathogenicity and Role of Plant Glutamine in Nitrogen-Induced Susceptibility (NIS) To Rice Blast
- PMID: 28293247
- PMCID: PMC5329020
- DOI: 10.3389/fpls.2017.00265
Increase of Fungal Pathogenicity and Role of Plant Glutamine in Nitrogen-Induced Susceptibility (NIS) To Rice Blast
Abstract
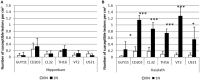
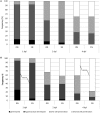
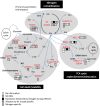
Highlight Modifications in glutamine synthetase OsGS1-2 expression and fungal pathogenicity underlie nitrogen-induced susceptibility to rice blast. Understanding why nitrogen fertilization increase the impact of many plant diseases is of major importance. The interaction between Magnaporthe oryzae and rice was used as a model for analyzing the molecular mechanisms underlying Nitrogen-Induced Susceptibility (NIS). We show that our experimental system in which nitrogen supply strongly affects rice blast susceptibility only slightly affects plant growth. In order to get insights into the mechanisms of NIS, we conducted a dual RNA-seq experiment on rice infected tissues under two nitrogen fertilization regimes. On the one hand, we show that enhanced susceptibility was visible despite an over-induction of defense gene expression by infection under high nitrogen regime. On the other hand, the fungus expressed to high levels effectors and pathogenicity-related genes in plants under high nitrogen regime. We propose that in plants supplied with elevated nitrogen fertilization, the observed enhanced induction of plant defense is over-passed by an increase in the expression of the fungal pathogenicity program, thus leading to enhanced susceptibility. Moreover, some rice genes implicated in nitrogen recycling were highly induced during NIS. We further demonstrate that the OsGS1-2 glutamine synthetase gene enhances plant resistance to M. oryzae and abolishes NIS and pinpoint glutamine as a potential key nutrient during NIS.
Keywords: Magnaporthe oryzae; defense; effector; fertilizer; glutamine; nitrogen; pathogenicity; rice.
Figures



Similar articles
-
Genome-wide association of rice response to blast fungus identifies loci for robust resistance under high nitrogen.BMC Plant Biol. 2021 Feb 18;21(1):99. doi: 10.1186/s12870-021-02864-3. BMC Plant Biol. 2021. PMID: 33602120 Free PMC article.
-
Phosphate accumulation in rice leaves promotes fungal pathogenicity and represses host immune responses during pathogen infection.Front Plant Sci. 2024 Jan 17;14:1330349. doi: 10.3389/fpls.2023.1330349. eCollection 2023. Front Plant Sci. 2024. PMID: 38298608 Free PMC article.
-
Diversity and genetics of nitrogen-induced susceptibility to the blast fungus in rice and wheat.Rice (N Y). 2013 Nov 20;6(1):32. doi: 10.1186/1939-8433-6-32. Rice (N Y). 2013. PMID: 24280346 Free PMC article.
-
Recent progress in understanding PAMP- and effector-triggered immunity against the rice blast fungus Magnaporthe oryzae.Mol Plant. 2013 May;6(3):605-20. doi: 10.1093/mp/sst015. Epub 2013 Jan 22. Mol Plant. 2013. PMID: 23340743 Review.
-
Phytohormones: the chemical language in Magnaporthe oryzae-rice pathosystem.Mycology. 2018 Jun 12;9(3):233-237. doi: 10.1080/21501203.2018.1483441. eCollection 2018. Mycology. 2018. PMID: 30181929 Free PMC article. Review.
Cited by
-
Evolutionary reconstruction, nomenclature and functional meta-analysis of the Kiwellin protein family.Front Plant Sci. 2022 Dec 22;13:1034708. doi: 10.3389/fpls.2022.1034708. eCollection 2022. Front Plant Sci. 2022. PMID: 36618657 Free PMC article.
-
Estimating Crop Nutritional Status Using Smart Apps to Support Nitrogen Fertilization. A Case Study on Paddy Rice.Sensors (Basel). 2019 Feb 25;19(4):981. doi: 10.3390/s19040981. Sensors (Basel). 2019. PMID: 30823623 Free PMC article.
-
Characterization of defense responses against bacterial pathogens in duckweeds lacking EDS1.New Phytol. 2022 Dec;236(5):1838-1855. doi: 10.1111/nph.18453. Epub 2022 Sep 25. New Phytol. 2022. PMID: 36052715 Free PMC article.
-
Multi-Omic Investigation of Low-Nitrogen Conditional Resistance to Clubroot Reveals Brassica napus Genes Involved in Nitrate Assimilation.Front Plant Sci. 2022 Feb 11;13:790563. doi: 10.3389/fpls.2022.790563. eCollection 2022. Front Plant Sci. 2022. PMID: 35222461 Free PMC article.
-
TaAMT2;3a, a wheat AMT2-type ammonium transporter, facilitates the infection of stripe rust fungus on wheat.BMC Plant Biol. 2019 Jun 6;19(1):239. doi: 10.1186/s12870-019-1841-8. BMC Plant Biol. 2019. PMID: 31170918 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

