Enzymatic improvement of mitochondrial thiol oxidase Erv1 for oxidized glutathione fermentation by Saccharomyces cerevisiae
- PMID: 28298220
- PMCID: PMC5353892
- DOI: 10.1186/s12934-017-0658-0
Enzymatic improvement of mitochondrial thiol oxidase Erv1 for oxidized glutathione fermentation by Saccharomyces cerevisiae
Abstract
Background: Oxidized glutathione (GSSG) is the preferred form for industrial mass production of glutathione due to its high stability compared with reduced glutathione (GSH). In our previous study, over-expression of the mitochondrial thiol oxidase ERV1 gene was the most effective for high GSSG production in Saccharomyces cerevisiae cells among three types of different thiol oxidase genes.
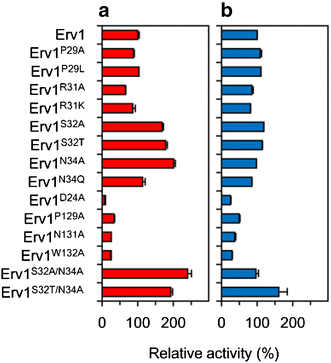
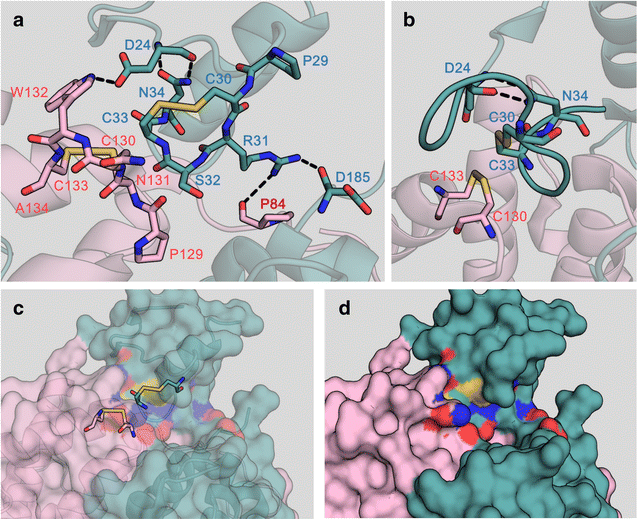
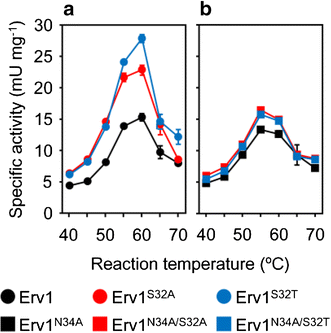
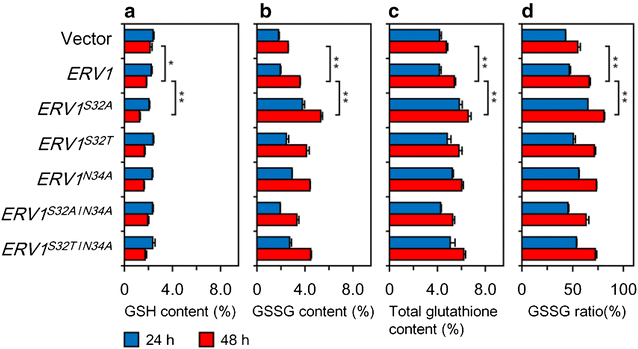
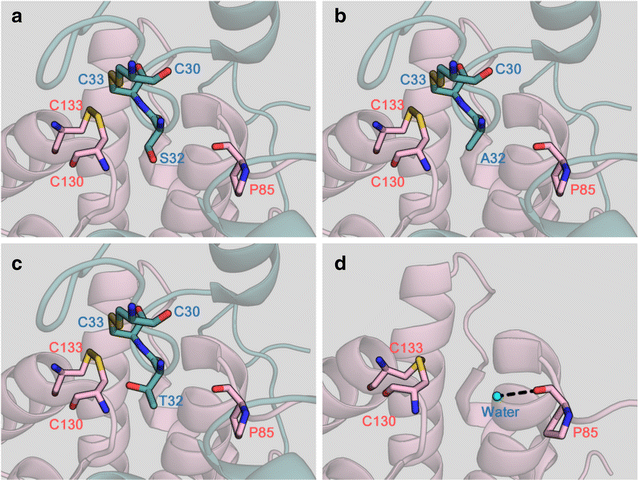
Results: We improved Erv1 enzyme activity for oxidation of GSH and revealed that S32 and N34 residues are critical for the oxidation. Five engineered Erv1 variant proteins containing S32 and/or N34 replacements exhibited 1.7- to 2.4-fold higher in vitro GSH oxidation activity than that of parental Erv1, whereas the oxidation activities of these variants for γ-glutamylcysteine were comparable. According to three-dimensional structures of Erv1 and protein stability assays, S32 and N34 residues interact with nearby residues through hydrogen bonding and greatly contribute to protein stability. These results suggest that increased flexibility by amino acid replacements around the active center decrease inhibitory effects on GSH oxidation. Over-expressions of mutant genes coding these Erv1 variants also increased GSSG and consequently total glutathione production in S. cerevisiae cells. Over-expression of the ERV1 S32A gene was the most effective for GSSG production in S. cerevisiae cells among the parent and other mutant genes, and it increased GSSG production about 1.5-fold compared to that of the parental ERV1 gene.
Conclusions: This is the first study demonstrating the pivotal effects of S32 and N34 residues to high GSH oxidation activity of Erv1. Furthermore, in vivo validity of Erv1 variants containing these S32 and N34 replacements were also demonstrated. This study indicates potentials of Erv1 for high GSSG production.
Keywords: Erv1; Glutathione; Mia40; Saccharomyces cerevisiae; Thiol oxidase.
Figures
Similar articles
-
Improvement of oxidized glutathione fermentation by thiol redox metabolism engineering in Saccharomyces cerevisiae.Appl Microbiol Biotechnol. 2015 Nov;99(22):9771-8. doi: 10.1007/s00253-015-6847-z. Epub 2015 Aug 4. Appl Microbiol Biotechnol. 2015. PMID: 26239069
-
Cytosolic Fe-S Cluster Protein Maturation and Iron Regulation Are Independent of the Mitochondrial Erv1/Mia40 Import System.J Biol Chem. 2015 Nov 13;290(46):27829-40. doi: 10.1074/jbc.M115.682179. Epub 2015 Sep 22. J Biol Chem. 2015. PMID: 26396185 Free PMC article.
-
Mitochondrial disulfide bond formation is driven by intersubunit electron transfer in Erv1 and proofread by glutathione.Mol Cell. 2010 Feb 26;37(4):516-28. doi: 10.1016/j.molcel.2010.01.017. Mol Cell. 2010. PMID: 20188670
-
Structural basis for the disulfide relay system in the mitochondrial intermembrane space.Antioxid Redox Signal. 2010 Nov 1;13(9):1359-73. doi: 10.1089/ars.2010.3099. Antioxid Redox Signal. 2010. PMID: 20136511 Review.
-
Glutathione production by Saccharomyces cerevisiae: current state and perspectives.Appl Microbiol Biotechnol. 2022 Mar;106(5-6):1879-1894. doi: 10.1007/s00253-022-11826-0. Epub 2022 Feb 19. Appl Microbiol Biotechnol. 2022. PMID: 35182192 Review.
Cited by
-
A Procedure for Precise Determination of Glutathione Produced by Saccharomyces cerevisiae.Bio Protoc. 2018 Jun 20;8(12):e2887. doi: 10.21769/BioProtoc.2887. eCollection 2018 Jun 20. Bio Protoc. 2018. PMID: 34285996 Free PMC article.
-
Metabolic engineering of the L-serine biosynthetic pathway improves glutathione production in Saccharomyces cerevisiae.Microb Cell Fact. 2022 Aug 6;21(1):153. doi: 10.1186/s12934-022-01880-8. Microb Cell Fact. 2022. PMID: 35933377 Free PMC article.
References
-
- Penninckx MJ. An overview on glutathione in Saccharomyces versus non-conventional yeasts. FEMS Yeast Res. 2002;2:295–305. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

