Evolutionary Analyses of GRAS Transcription Factors in Angiosperms
- PMID: 28303145
- PMCID: PMC5332381
- DOI: 10.3389/fpls.2017.00273
Evolutionary Analyses of GRAS Transcription Factors in Angiosperms
Abstract
GRAS transcription factors (TFs) play critical roles in plant growth and development such as gibberellin and mycorrhizal signaling. Proteins belonging to this gene family contain a typical GRAS domain in the C-terminal sequence, whereas the N-terminal region is highly variable. Although, GRAS genes have been characterized in a number of plant species, their classification is still not completely resolved. Based on a panel of eight representative species of angiosperms, we identified 29 orthologous groups or orthogroups (OGs) for the GRAS gene family, suggesting that at least 29 ancestor genes were present in the angiosperm lineage before the "Amborella" evolutionary split. Interestingly, some taxonomic groups were missing members of one or more OGs. The gene number expansion usually observed in transcription factors was not observed in GRAS while the genome triplication ancestral to the eudicots (γ hexaploidization event) was detectable in a limited number of GRAS orthogroups. We also found conserved OG-specific motifs in the variable N-terminal region. Finally, we could regroup OGs in 17 subfamilies for which names were homogenized based on a literature review and described 5 new subfamilies (DLT, RAD1, RAM1, SCLA, and SCLB). This study establishes a consistent framework for the classification of GRAS members in angiosperm species, and thereby a tool to correctly establish the orthologous relationships of GRAS genes in most of the food crops in order to facilitate any subsequent functional analyses in the GRAS gene family. The multi-fasta file containing all the sequences used in our study could be used as database to perform diagnostic BLASTp to classify GRAS genes from other non-model species.
Keywords: GRAS gene family; plant evolution; transcription factors.
Figures
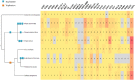


References
-
- Abarca D., Pizarro A., Hernández I., Sánchez C., Solana S. P., Del Amo A., et al. . (2014). The GRAS gene family in pine: transcript expression patterns associated with the maturation-related decline of competence to form adventitious roots. BMC Plant Biol. 14:354. 10.1186/s12870-014-0354-8 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

