IsoSel: Protein Isoform Selector for phylogenetic reconstructions
- PMID: 28323858
- PMCID: PMC5360266
- DOI: 10.1371/journal.pone.0174250
IsoSel: Protein Isoform Selector for phylogenetic reconstructions
Abstract
The reliability of molecular phylogenies is strongly dependent on the quality of the assembled datasets. In the case of eukaryotes, the selection of only one protein isoform per genomic locus is mandatory to avoid biases linked to redundancy. Here, we present IsoSel, a tool devoted to the selection of alternative isoforms in the context of phylogenetic reconstruction. It provides a better alternative to the widely used approach consisting in the selection of the longest isoforms and it performs better than Guidance, its only available counterpart. IsoSel is publicly available at http://doua.prabi.fr/software/isosel.
Conflict of interest statement
Figures
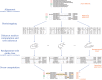
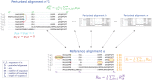
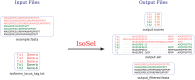



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

