Neuropathic pain promotes adaptive changes in gene expression in brain networks involved in stress and depression
- PMID: 28325815
- PMCID: PMC5524975
- DOI: 10.1126/scisignal.aaj1549
Neuropathic pain promotes adaptive changes in gene expression in brain networks involved in stress and depression
Abstract
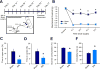
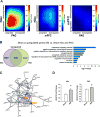
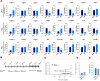
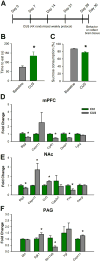
Neuropathic pain is a complex chronic condition characterized by various sensory, cognitive, and affective symptoms. A large percentage of patients with neuropathic pain are also afflicted with depression and anxiety disorders, a pattern that is also seen in animal models. Furthermore, clinical and preclinical studies indicate that chronic pain corresponds with adaptations in several brain networks involved in mood, motivation, and reward. Chronic stress is also a major risk factor for depression. We investigated whether chronic pain and stress affect similar molecular mechanisms and whether chronic pain can affect gene expression patterns that are involved in depression. Using two mouse models of neuropathic pain and depression [spared nerve injury (SNI) and chronic unpredictable stress (CUS)], we performed next-generation RNA sequencing and pathway analysis to monitor changes in gene expression in the nucleus accumbens (NAc), the medial prefrontal cortex (mPFC), and the periaqueductal gray (PAG). In addition to finding unique transcriptome profiles across these regions, we identified a substantial number of signaling pathway-associated genes with similar changes in expression in both SNI and CUS mice. Many of these genes have been implicated in depression, anxiety, and chronic pain in patients. Our study provides a resource of the changes in gene expression induced by long-term neuropathic pain in three distinct brain regions and reveals molecular connections between pain and chronic stress.
Copyright © 2017, American Association for the Advancement of Science.
Conflict of interest statement
Figures



References
-
- C. Committee on Advancing Pain Research, and Education. Relieving Pain in America: A Blueprint for Transforming Prevention, Care, Education, and Research. Institute of Medicine (US); Washington, DC: 2011.
-
- Lerman SF, Rudich Z, Brill S, Shalev H, Shahar G. Longitudinal associations between depression, anxiety, pain, and pain-related disability in chronic pain patients. Psychosomatic medicine. 2015;77:333–341. - PubMed
-
- McWilliams LA, Goodwin RD, Cox BJ. Depression and anxiety associated with three pain conditions: results from a nationally representative sample. Pain. 2004;111:77–83. - PubMed
-
- Taylor AM, Castonguay A, Taylor AJ, Murphy NP, Ghogha A, Cook C, Xue L, Olmstead MC, De Koninck Y, Evans CJ, Cahill CM. Microglia disrupt mesolimbic reward circuitry in chronic pain. The Journal of neuroscience : the official journal of the Society for Neuroscience. 2015;35:8442–8450. - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

