Dual-utility NLS drives RNF169-dependent DNA damage responses
- PMID: 28325877
- PMCID: PMC5389323
- DOI: 10.1073/pnas.1616602114
Dual-utility NLS drives RNF169-dependent DNA damage responses
Abstract
Loading of p53-binding protein 1 (53BP1) and receptor-associated protein 80 (RAP80) at DNA double-strand breaks (DSBs) drives cell cycle checkpoint activation but is counterproductive to high-fidelity DNA repair. ring finger protein 169 (RNF169) maintains the balance by limiting the deposition of DNA damage mediator proteins at the damaged chromatin. We report here that this attribute is accomplished, in part, by a predicted nuclear localization signal (NLS) that not only shuttles RNF169 into the nucleus but also promotes its stability by mediating a direct interaction with the ubiquitin-specific protease USP7. Guided by the crystal structure of USP7 in complex with the RNF169 NLS, we uncoupled USP7 binding from its nuclear import function and showed that perturbing the USP7-RNF169 complex destabilized RNF169, compromised high-fidelity DSB repair, and hypersensitized cells to poly (ADP-ribose) polymerase inhibition. Finally, expression of USP7 and RNF169 positively correlated in breast cancer specimens. Collectively, our findings uncover an NLS-mediated bipartite mechanism that supports the nuclear function of a DSB response protein.
Keywords: DNA damage; DNA repair; RNF169; USP7; deubiquitylation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




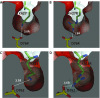



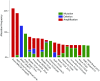
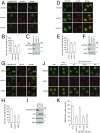




Comment in
-
NLS protects DNA damage protein.Sci Signal. 2017 Apr 11;10(474):eaan4046. doi: 10.1126/scisignal.aan4046. Sci Signal. 2017. PMID: 28400536
References
-
- Schwertman P, Bekker-Jensen S, Mailand N. Regulation of DNA double-strand break repair by ubiquitin and ubiquitin-like modifiers. Nat Rev Mol Cell Biol. 2016;17(6):379–394. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

