A sensitivity analysis of RNA folding nearest neighbor parameters identifies a subset of free energy parameters with the greatest impact on RNA secondary structure prediction
- PMID: 28334976
- PMCID: PMC5449625
- DOI: 10.1093/nar/gkx170
A sensitivity analysis of RNA folding nearest neighbor parameters identifies a subset of free energy parameters with the greatest impact on RNA secondary structure prediction
Abstract
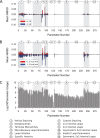
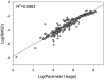
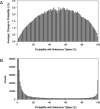
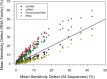
Nearest neighbor parameters for estimating the folding energy changes of RNA secondary structures are used in structure prediction and analysis. Despite their widespread application, a comprehensive analysis of the impact of each parameter on the precision of calculations had not been conducted. To identify the parameters with greatest impact, a sensitivity analysis was performed on the 291 parameters that compose the 2004 version of the free energy nearest neighbor rules. Perturbed parameter sets were generated by perturbing each parameter independently. Then the effect of each individual parameter change on predicted base-pair probabilities and secondary structures as compared to the standard parameter set was observed for a set of sequences including structured ncRNA, mRNA and randomized sequences. The results identify for the first time the parameters with the greatest impact on secondary structure prediction, and the subset which should be prioritized for further study in order to improve the precision of structure prediction. In particular, bulge loop initiation, multibranch loop initiation, AU/GU internal loop closure and AU/GU helix end parameters were particularly important. An analysis of parameter usage during folding free energy calculations of stochastic samples of secondary structures revealed a correlation between parameter usage and impact on structure prediction precision.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
References
-
- Wu L., Belasco J.G.. Let me count the ways: mechanisms of gene regulation by miRNAs and siRNAs. Mol. Cell. 2008; 29:1–7. - PubMed
-
- Doudna J.A., Cech T.R.. The chemical repertoire of natural ribozymes. Nature. 2002; 418:222–228. - PubMed
-
- Tinoco I. Jr, Bustamante C.. How RNA folds. J. Mol. Biol. 1999; 293:271–281. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

