CancerLocator: non-invasive cancer diagnosis and tissue-of-origin prediction using methylation profiles of cell-free DNA
- PMID: 28335812
- PMCID: PMC5364586
- DOI: 10.1186/s13059-017-1191-5
CancerLocator: non-invasive cancer diagnosis and tissue-of-origin prediction using methylation profiles of cell-free DNA
Abstract
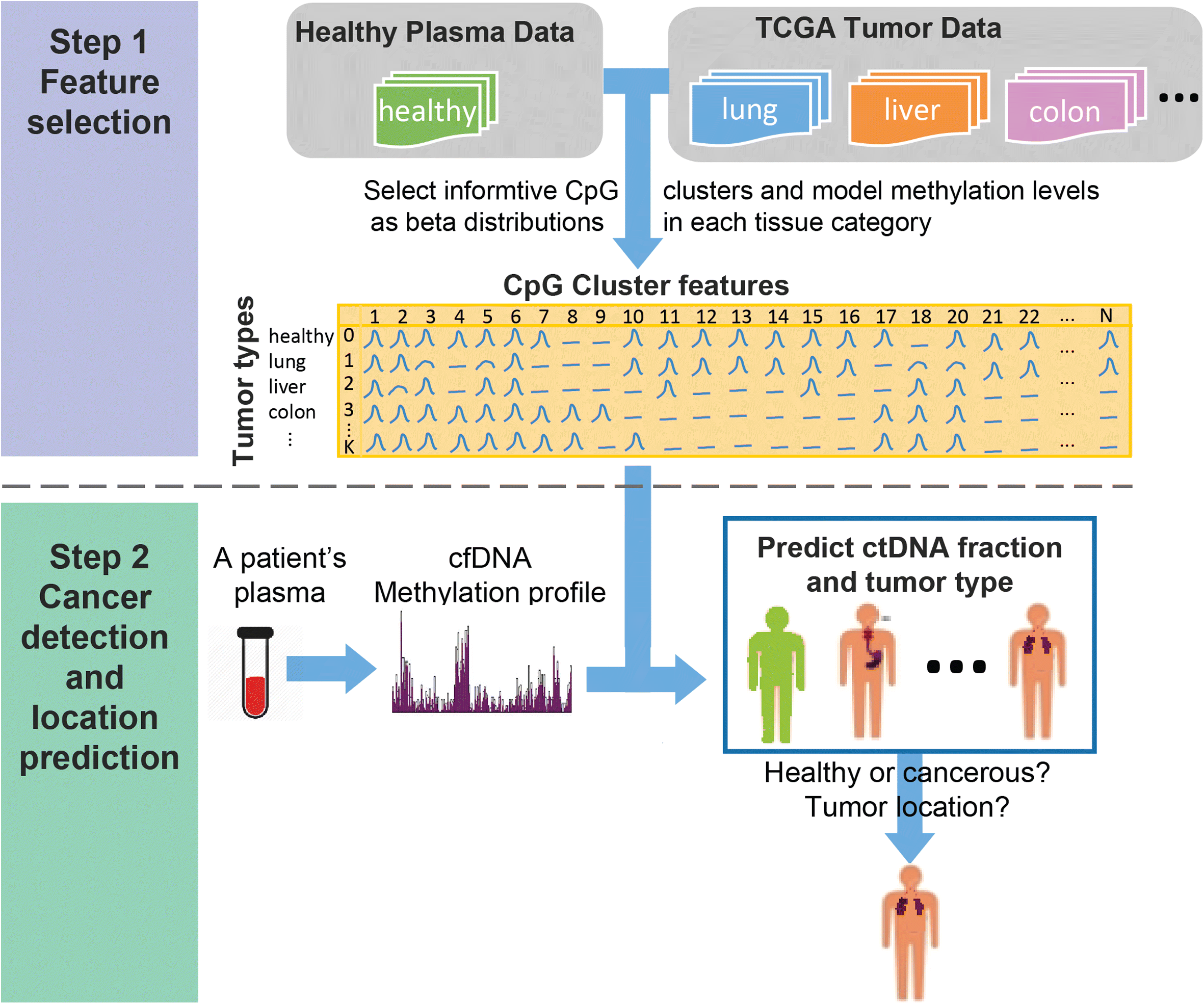
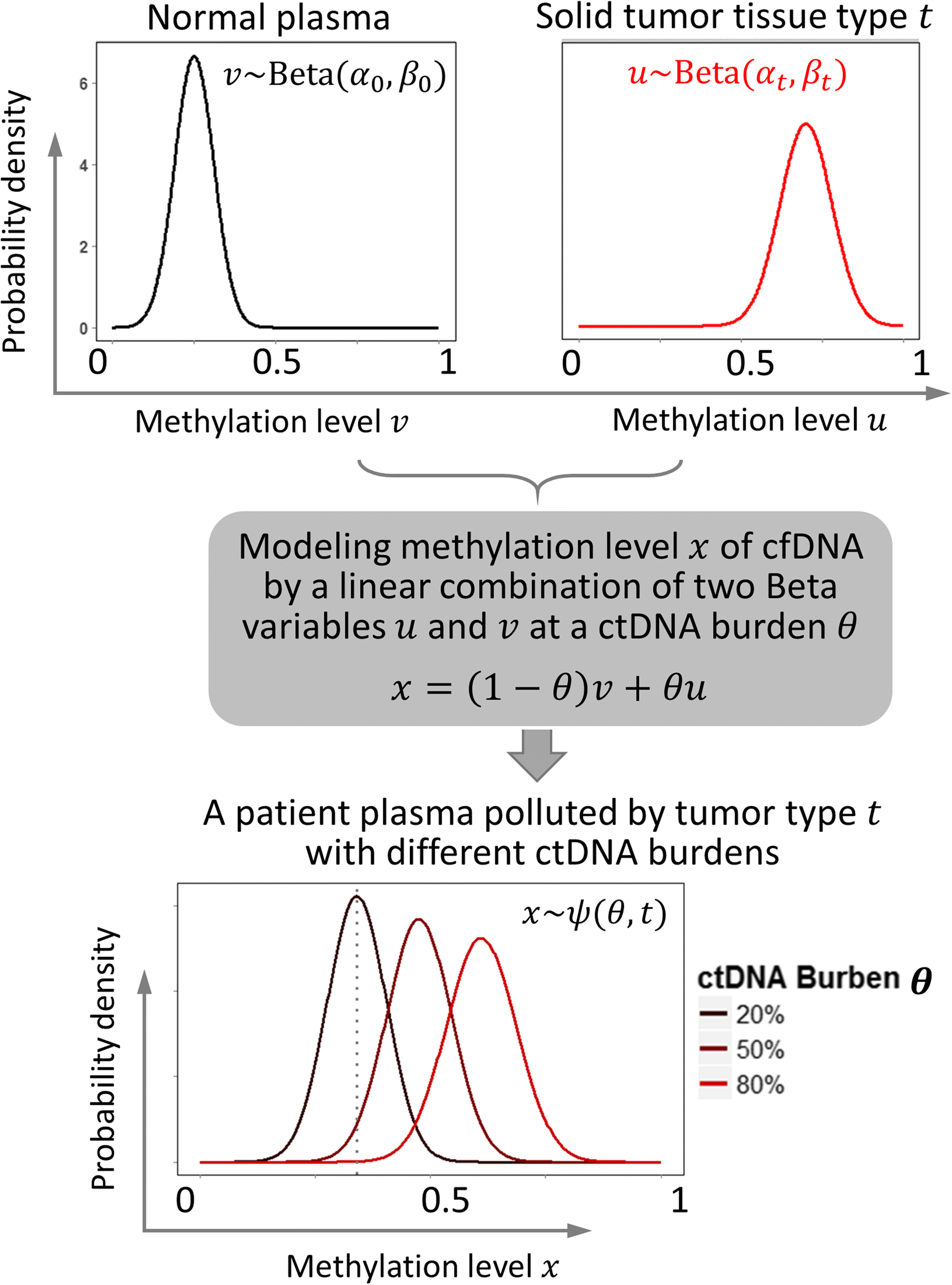
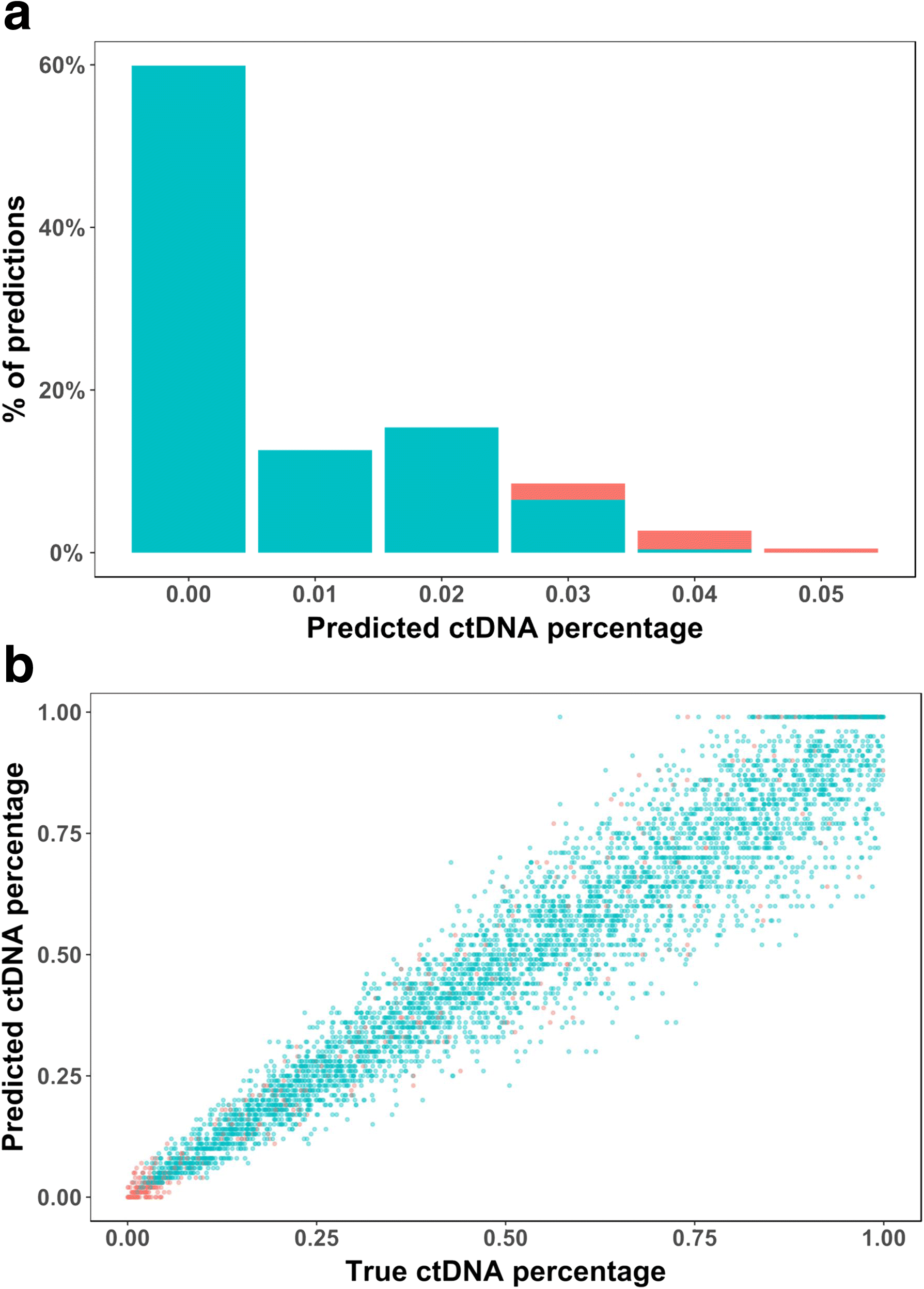
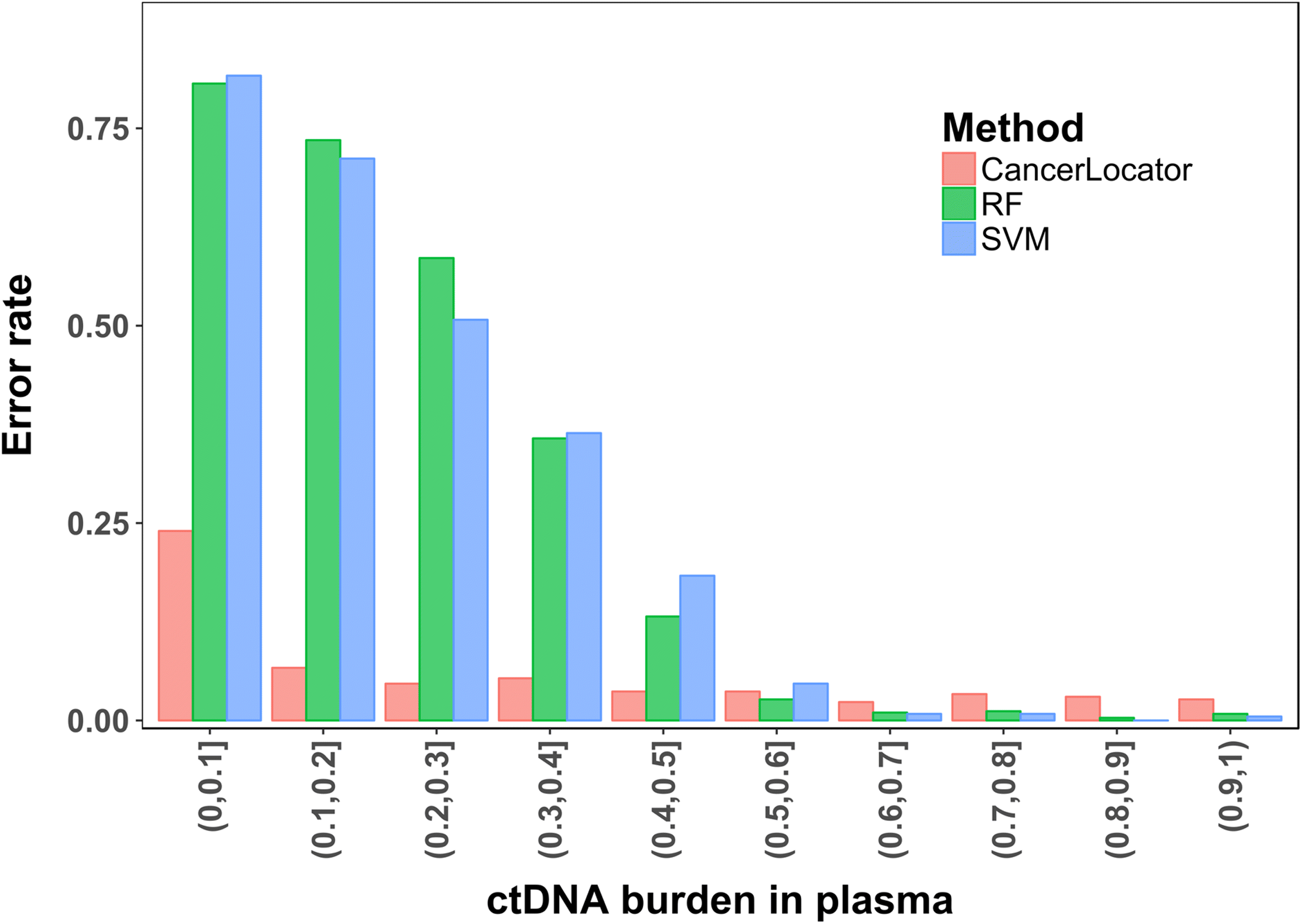
We propose a probabilistic method, CancerLocator, which exploits the diagnostic potential of cell-free DNA by determining not only the presence but also the location of tumors. CancerLocator simultaneously infers the proportions and the tissue-of-origin of tumor-derived cell-free DNA in a blood sample using genome-wide DNA methylation data. CancerLocator outperforms two established multi-class classification methods on simulations and real data, even with the low proportion of tumor-derived DNA in the cell-free DNA scenarios. CancerLocator also achieves promising results on patient plasma samples with low DNA methylation sequencing coverage.
Keywords: Cancer diagnosis; Cell-free DNA; DNA methylation; Liquid biopsy; Next-generation sequencing.
Figures