Subcellular localization of Arabidopsis arogenate dehydratases suggests novel and non-enzymatic roles
- PMID: 28338876
- PMCID: PMC5444438
- DOI: 10.1093/jxb/erx024
Subcellular localization of Arabidopsis arogenate dehydratases suggests novel and non-enzymatic roles
Abstract
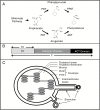
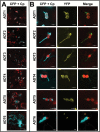
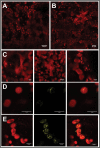
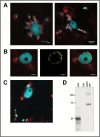
Arogenate dehydratases (ADTs) catalyze the final step in phenylalanine biosynthesis in plants. The Arabidopsis thaliana genome encodes a family of six ADTs capable of decarboxylating/dehydrating arogenate into phenylalanine. Using cyan fluorescent protein (CFP)-tagged proteins, the subcellular localization patterns of all six A. thaliana ADTs were investigated in intact Nicotiana benthamiana and A. thaliana leaf cells. We show that A. thaliana ADTs localize to stroma and stromules (stroma-filled tubules) of chloroplasts. This localization pattern is consistent with the enzymatic function of ADTs as many enzymes required for amino acid biosynthesis are primarily localized to chloroplasts, and stromules are thought to increase metabolite transport from chloroplasts to other cellular compartments. Furthermore, we provide evidence that ADTs have additional, non-enzymatic roles. ADT2 localizes in a ring around the equatorial plane of chloroplasts or to a chloroplast pole, which suggests that ADT2 is a component of the chloroplast division machinery. In addition to chloroplasts, ADT5 was also found in nuclei, again suggesting a non-enzymatic role for ADT5. We also show evidence that ADT5 is transported to the nucleus via stromules. We propose that ADT2 and ADT5 are moonlighting proteins that play an enzymatic role in phenylalanine biosynthesis and a second role in chloroplast division or transcriptional regulation, respectively.
Keywords: Arogenate dehydratase; chloroplast division; moonlighting proteins; nuclear localization; phenylalanine biosynthesis; stromules..
© The Author 2017. Published by Oxford University Press on behalf of the Society for Experimental Biology.
Figures



References
-
- Agati G, Biricolti S, Guidi L, Ferrini F, Fini A, Tattini M. 2011. The biosynthesis of flavonoids is enhanced similarly by UV radiation and root zone salinity in L. vulgare leaves. Journal of Plant Physiology 168, 204–212. - PubMed
-
- Bross CD, Corea ORA, Kaldis A, Menassa R, Bernards MA, Kohalmi SE. 2011. Complementation of the pha2 yeast mutant suggests functional differences for arogenate dehydratases from Arabidopsis thaliana. Plant Physiology and Biochemistry 49, 882–890. - PubMed
-
- Cho MH, Corea ORA, Yang H, et al. 2007. Phenylalanine biosynthesis in Arabidopsis thaliana. Identification and characterization of arogenate dehydratases. Journal of Biological Chemistry 282, 30827–30835. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

