Genome-Wide Estimates of Transposable Element Insertion and Deletion Rates in Drosophila Melanogaster
- PMID: 28338986
- PMCID: PMC5447328
- DOI: 10.1093/gbe/evx050
Genome-Wide Estimates of Transposable Element Insertion and Deletion Rates in Drosophila Melanogaster
Abstract
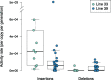
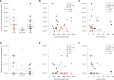
Knowing the rate at which transposable elements (TEs) insert and delete is critical for understanding their role in genome evolution. We estimated spontaneous rates of insertion and deletion for all known, active TE superfamilies present in a set of Drosophila melanogaster mutation-accumulation (MA) lines using whole genome sequence data. Our results demonstrate that TE insertions far outpace TE deletions in D. melanogaster. We found a significant effect of background genotype on TE activity, with higher rates of insertions in one MA line. We also found significant rate heterogeneity between the chromosomes, with both insertion and deletion rates elevated on the X relative to the autosomes. Further, we identified significant associations between TE activity and chromatin state, and tested for associations between TE activity and other features of the local genomic environment such as TE content, exon content, GC content, and recombination rate. Our results provide the most detailed assessment of TE mobility in any organism to date, and provide a useful benchmark for both addressing theoretical predictions of TE dynamics and for exploring large-scale patterns of TE movement in D. melanogaster and other species.
Keywords: Drosophila melanogaster; deletion rate; insertion rate; transposable elements; transposition.
© The Author(s) 2017. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures

References
-
- Bartolom� C, Maside X, Charlesworth B. 2002. On the abundance and distribution of transposable elements in the genome of Drosophila melanogaster. Mol Biol Evol. 19:926–937. - PubMed
-
- Brennecke J, et al. 2007. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128:1089–1103. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

