PML protein organizes heterochromatin domains where it regulates histone H3.3 deposition by ATRX/DAXX
- PMID: 28341773
- PMCID: PMC5453325
- DOI: 10.1101/gr.215830.116
PML protein organizes heterochromatin domains where it regulates histone H3.3 deposition by ATRX/DAXX
Abstract
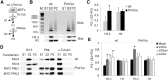
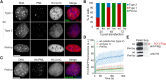
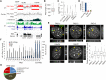
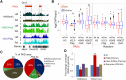
Maintenance of chromatin homeostasis involves proper delivery of histone variants to the genome. The interplay between different chaperones regulating the supply of histone variants to distinct chromatin domains remains largely undeciphered. We report a role of promyelocytic leukemia (PML) protein in the routing of histone variant H3.3 to chromatin and in the organization of megabase-size heterochromatic PML-associated domains that we call PADs. Loss of PML alters the heterochromatic state of PADs by shifting the histone H3 methylation balance from K9me3 to K27me3. Loss of PML impairs deposition of H3.3 by ATRX and DAXX in PADs but preserves the H3.3 loading function of HIRA in these regions. Our results unveil an unappreciated role of PML in the large-scale organization of chromatin and demonstrate a PML-dependent role of ATRX/DAXX in the deposition of H3.3 in PADs. Our data suggest that H3.3 loading by HIRA and ATRX-dependent H3K27 trimethylation constitute mechanisms ensuring maintenance of heterochromatin when the integrity of these domains is compromised.
© 2017 Delbarre et al.; Published by Cold Spring Harbor Laboratory Press.
Figures
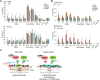
Similar articles
-
PML is recruited to heterochromatin during S phase and represses DAXX-mediated histone H3.3 chromatin assembly.J Cell Sci. 2019 Mar 26;132(6):jcs220970. doi: 10.1242/jcs.220970. J Cell Sci. 2019. PMID: 30796101 Free PMC article.
-
Long non-coding RNA ChRO1 facilitates ATRX/DAXX-dependent H3.3 deposition for transcription-associated heterochromatin reorganization.Nucleic Acids Res. 2018 Dec 14;46(22):11759-11775. doi: 10.1093/nar/gky923. Nucleic Acids Res. 2018. PMID: 30335163 Free PMC article.
-
Dynamics of histone H3.3 deposition in proliferating and senescent cells reveals a DAXX-dependent targeting to PML-NBs important for pericentromeric heterochromatin organization.Cell Cycle. 2014;13(2):249-67. doi: 10.4161/cc.26988. Epub 2013 Nov 5. Cell Cycle. 2014. PMID: 24200965 Free PMC article.
-
Modulation of H3.3 chromatin assembly by PML: A way to regulate epigenetic inheritance.Bioessays. 2021 Oct;43(10):e2100038. doi: 10.1002/bies.202100038. Epub 2021 Aug 22. Bioessays. 2021. PMID: 34423467 Review.
-
New players in heterochromatin silencing: histone variant H3.3 and the ATRX/DAXX chaperone.Nucleic Acids Res. 2016 Feb 29;44(4):1496-501. doi: 10.1093/nar/gkw012. Epub 2016 Jan 14. Nucleic Acids Res. 2016. PMID: 26773061 Free PMC article. Review.
Cited by
-
PML Nuclear bodies: the cancer connection and beyond.Nucleus. 2024 Dec;15(1):2321265. doi: 10.1080/19491034.2024.2321265. Epub 2024 Feb 27. Nucleus. 2024. PMID: 38411156 Free PMC article. Review.
-
PML-mediated nuclear loosening permits immunomodulation of mesenchymal stem/stromal cells under inflammatory conditions.Cell Prolif. 2024 Apr;57(4):e13566. doi: 10.1111/cpr.13566. Epub 2023 Oct 20. Cell Prolif. 2024. PMID: 37864298 Free PMC article.
-
The Multiple Facets of ATRX Protein.Cancers (Basel). 2021 May 5;13(9):2211. doi: 10.3390/cancers13092211. Cancers (Basel). 2021. PMID: 34062956 Free PMC article. Review.
-
Rapid and reversible suppression of ALT by DAXX in osteosarcoma cells.Sci Rep. 2019 Mar 14;9(1):4544. doi: 10.1038/s41598-019-41058-8. Sci Rep. 2019. PMID: 30872698 Free PMC article.
-
Mechanistic dissection of dominant AIRE mutations in mouse models reveals AIRE autoregulation.J Exp Med. 2021 Nov 1;218(11):e20201076. doi: 10.1084/jem.20201076. Epub 2021 Sep 3. J Exp Med. 2021. PMID: 34477806 Free PMC article.
References
-
- Adam S, Polo SE, Almouzni G. 2013. Transcription recovery after DNA damage requires chromatin priming by the H3.3 histone chaperone HIRA. Cell 155: 94–106. - PubMed
-
- Bernardi R, Pandolfi PP. 2007. Structure, dynamics and functions of promyelocytic leukaemia nuclear bodies. Nat Rev Mol Cell Biol 8: 1006–1016. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
