Herboxidiene triggers splicing repression and abiotic stress responses in plants
- PMID: 28347276
- PMCID: PMC5369228
- DOI: 10.1186/s12864-017-3656-z
Herboxidiene triggers splicing repression and abiotic stress responses in plants
Abstract
Background: Constitutive and alternative splicing of pre-mRNAs from multiexonic genes controls the diversity of the proteome; these precisely regulated processes also fine-tune responses to cues related to growth, development, and stresses. Small-molecule inhibitors that perturb splicing provide invaluable tools for use as chemical probes to uncover the molecular underpinnings of splicing regulation and as potential anticancer compounds.
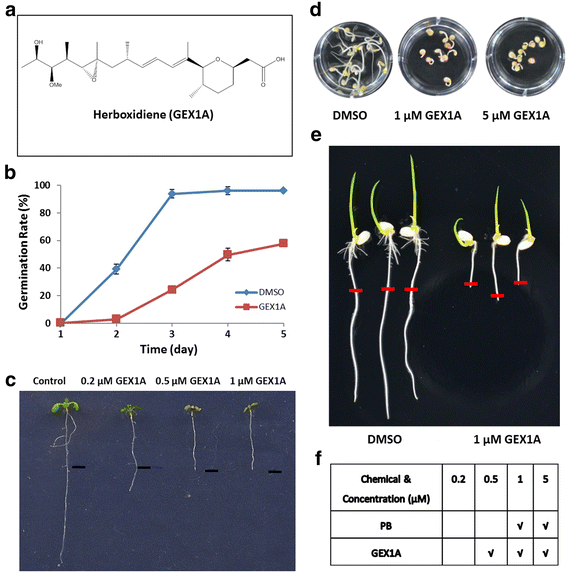
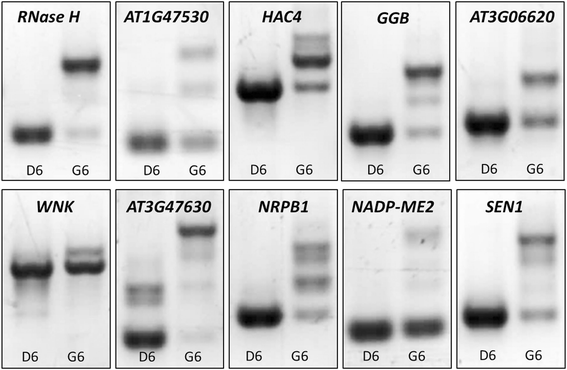
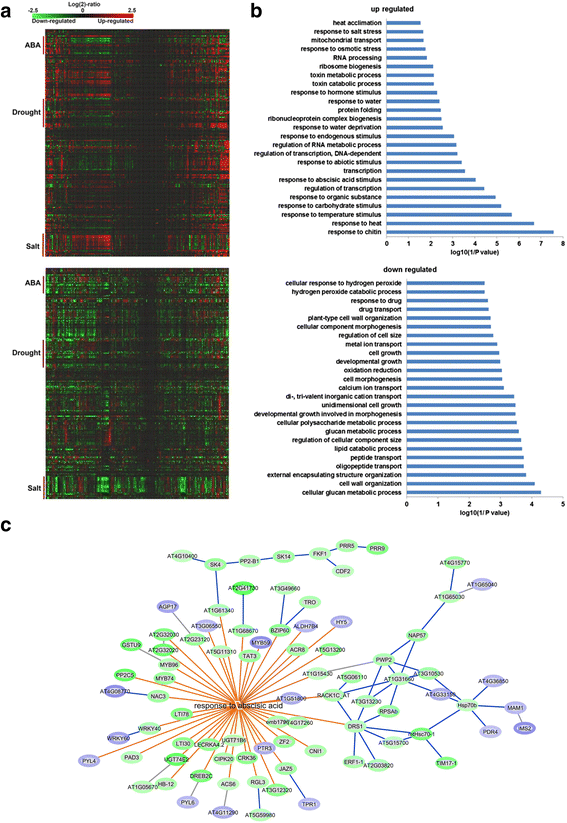
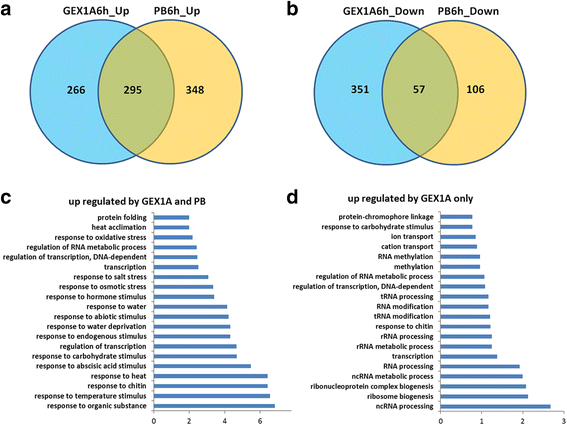
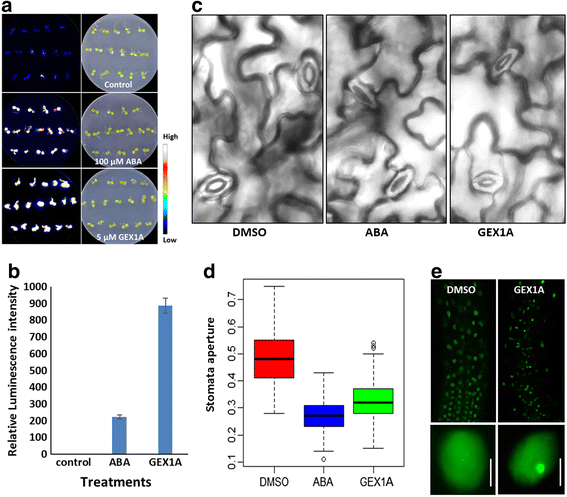
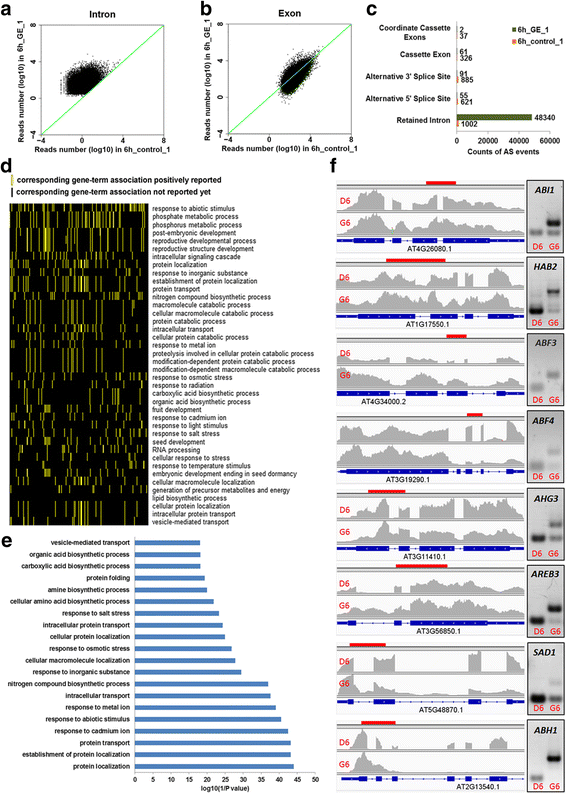
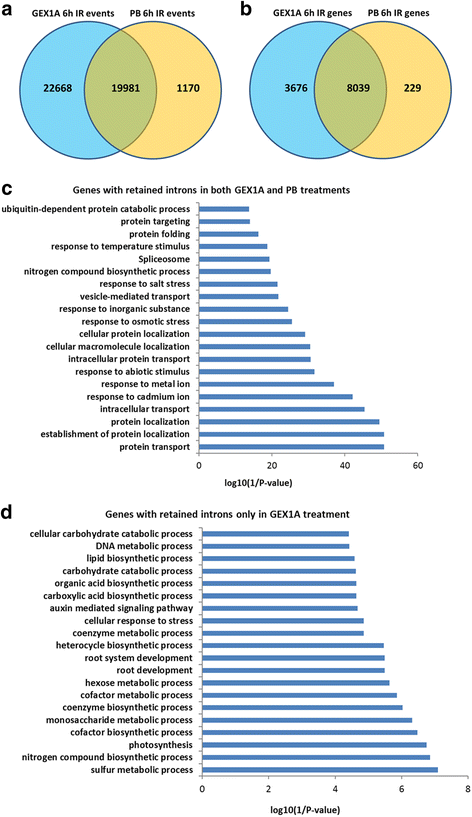
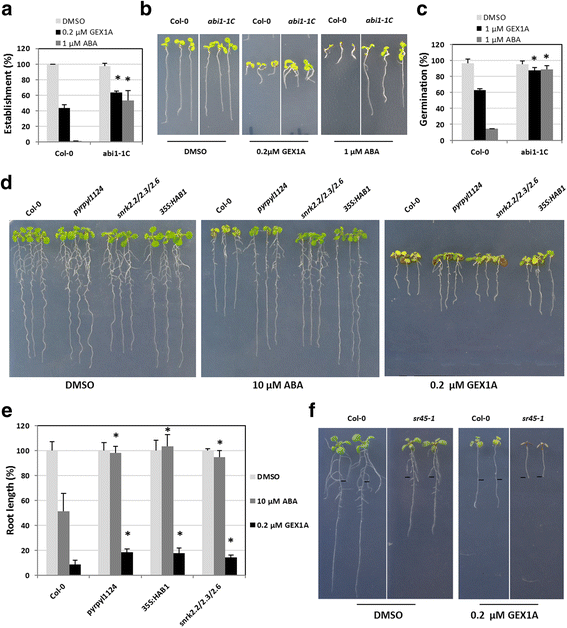
Results: Here, we show that herboxidiene (GEX1A) inhibits both constitutive and alternative splicing. Moreover, GEX1A activates genome-wide transcriptional patterns involved in abiotic stress responses in plants. GEX1A treatment -activated ABA-inducible promoters, and led to stomatal closure. Interestingly, GEX1A and pladienolide B (PB) elicited similar cellular changes, including alterations in the patterns of transcription and splicing, suggesting that these compounds might target the same spliceosome complex in plant cells.
Conclusions: Our study establishes GEX1A as a potent splicing inhibitor in plants that can be used to probe the assembly, dynamics, and molecular functions of the spliceosome and to study the interplay between splicing stress and abiotic stresses, as well as having potential biotechnological applications.
Keywords: ABA; Abiotic stress responses; Alternative splicing; GEX1A; Pladienolide B; SR proteins; Splicing inhibitors.
Figures
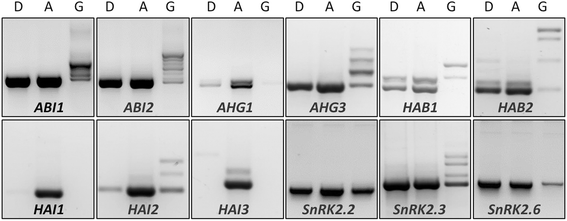
Similar articles
-
Pre-mRNA splicing repression triggers abiotic stress signaling in plants.Plant J. 2017 Jan;89(2):291-309. doi: 10.1111/tpj.13383. Epub 2017 Jan 17. Plant J. 2017. PMID: 27664942
-
Plant serine/arginine-rich proteins: roles in precursor messenger RNA splicing, plant development, and stress responses.Wiley Interdiscip Rev RNA. 2011 Nov-Dec;2(6):875-89. doi: 10.1002/wrna.98. Epub 2011 Jul 15. Wiley Interdiscip Rev RNA. 2011. PMID: 21766458 Review.
-
A homolog of splicing factor SF1 is essential for development and is involved in the alternative splicing of pre-mRNA in Arabidopsis thaliana.Plant J. 2014 May;78(4):591-603. doi: 10.1111/tpj.12491. Epub 2014 Apr 23. Plant J. 2014. PMID: 24580679
-
Identification of SAP155 as the target of GEX1A (Herboxidiene), an antitumor natural product.ACS Chem Biol. 2011 Mar 18;6(3):229-33. doi: 10.1021/cb100248e. Epub 2011 Jan 13. ACS Chem Biol. 2011. PMID: 21138297
-
Regulation of the circadian clock through pre-mRNA splicing in Arabidopsis.J Exp Bot. 2014 May;65(8):1973-80. doi: 10.1093/jxb/eru085. Epub 2014 Mar 6. J Exp Bot. 2014. PMID: 24604736 Review.
Cited by
-
Phosphorylation mediated regulation of RNA splicing in plants.Front Plant Sci. 2023 Sep 14;14:1249057. doi: 10.3389/fpls.2023.1249057. eCollection 2023. Front Plant Sci. 2023. PMID: 37780493 Free PMC article. Review.
-
A defective splicing machinery promotes senescence through MDM4 alternative splicing.Aging Cell. 2024 Nov;23(11):e14301. doi: 10.1111/acel.14301. Epub 2024 Aug 8. Aging Cell. 2024. PMID: 39118304 Free PMC article.
-
The U1 snRNP Subunit LUC7 Modulates Plant Development and Stress Responses via Regulation of Alternative Splicing.Plant Cell. 2018 Nov;30(11):2838-2854. doi: 10.1105/tpc.18.00244. Epub 2018 Oct 11. Plant Cell. 2018. PMID: 30309899 Free PMC article.
-
Multiplex CRISPR Mutagenesis of the Serine/Arginine-Rich (SR) Gene Family in Rice.Genes (Basel). 2019 Aug 7;10(8):596. doi: 10.3390/genes10080596. Genes (Basel). 2019. PMID: 31394891 Free PMC article.
-
Plant snRNP Biogenesis: A Perspective from the Nucleolus and Cajal Bodies.Front Plant Sci. 2018 Jan 4;8:2184. doi: 10.3389/fpls.2017.02184. eCollection 2017. Front Plant Sci. 2018. PMID: 29354141 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

