Helminth-induced alterations of the gut microbiota exacerbate bacterial colitis
- PMID: 28352104
- PMCID: PMC5620113
- DOI: 10.1038/mi.2017.20
Helminth-induced alterations of the gut microbiota exacerbate bacterial colitis
Abstract
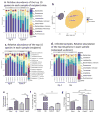
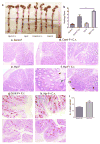
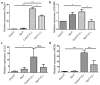
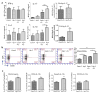
Infection with the intestinal helminth parasite Heligmosomoides polygyrus exacerbates the colitis caused by the bacterial enteropathogen Citrobacter rodentium. To clarify the underlying mechanism, we analyzed fecal microbiota composition of control and helminth-infected mice and evaluated the functional role of compositional differences by microbiota transplantation experiments. Our results showed that infection of Balb/c mice with H. polygyrus resulted in significant changes in the composition of the gut microbiota, characterized by a marked increase in the abundance of Bacteroidetes and decreases in Firmicutes and Lactobacillales. Recipients of the gut microbiota from helminth-infected wide-type, but not STAT6-deficient, Balb/c donors had increased fecal pathogen shedding and significant worsening of Citrobacter-induced colitis compared to recipients of microbiota from control donors. Recipients of helminth-altered microbiota also displayed increased regulatory T cells and IL-10 expression. Depletion of CD4+CD25+ T cells and neutralization of IL-10 in recipients of helminth-altered microbiota led to reduced stool C. rodentium numbers and attenuated colitis. These results indicate that alteration of the gut microbiota is a significant contributor to the H. polygyrus-induced exacerbation of C. rodentium colitis. The helminth-induced alteration of the microbiota is Th2-dependent and acts by promoting regulatory T cells that suppress protective responses to bacterial enteropathogens.
Conflict of interest statement
The authors have no conflict of interest to declare.
Figures



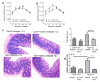
References
-
- Bäckhed F, Ley R, Sonnenburg J, Peterson D, Gordon J. Host-bacterial mutualism in the human intestine. Science (New York, NY) 2005;307(5717):1915–1920. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

