A Novel Genetic Screen Identifies Modifiers of Age-Dependent Amyloid β Toxicity in the Drosophila Brain
- PMID: 28352227
- PMCID: PMC5349081
- DOI: 10.3389/fnagi.2017.00061
A Novel Genetic Screen Identifies Modifiers of Age-Dependent Amyloid β Toxicity in the Drosophila Brain
Abstract
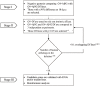
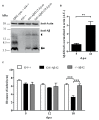
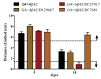
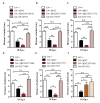
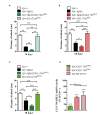
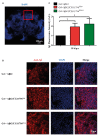
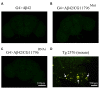
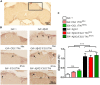
The accumulation of amyloid β peptide (Aβ) in the brain of Alzheimer's disease (AD) patients begins many years before clinical onset. Such process has been proposed to be pathogenic through the toxicity of Aβ soluble oligomers leading to synaptic dysfunction, phospho-tau aggregation and neuronal loss. Yet, a massive accumulation of Aβ can be found in approximately 30% of aged individuals with preserved cognitive function. Therefore, within the frame of the "amyloid hypothesis", compensatory mechanisms and/or additional neurotoxic or protective factors need to be considered and investigated. Here we describe a modifier genetic screen in Drosophila designed to identify genes that modulate toxicity of Aβ42 in the CNS. The expression of Aβ42 led to its accumulation in the brain and a moderate impairment of negative geotaxis at 18 days post-eclosion (d.p.e) as compared with genetic or parental controls. These flies were mated with a collection of lines carrying chromosomal deletions and negative geotaxis was assessed at 5 and 18 d.p.e. Our screen is the first to take into account all of the following features, relevant to sporadic AD: (1) pan-neuronal expression of wild-type Aβ42; (2) a quantifiable complex behavior; (3) Aβ neurotoxicity associated with progressive accumulation of the peptide; and (4) improvement or worsening of climbing ability only evident in aged animals. One hundred and ninety-nine deficiency (Df) lines accounting for ~6300 genes were analyzed. Six lines, including the deletion of 52 Drosophila genes with human orthologs, significantly modified Aβ42 neurotoxicity in 18-day-old flies. So far, we have validated CG11796 and identified CG17249 as a strong candidate (whose human orthologs are HPD and PRCC, respectively) by using RNAi or mutant hemizygous lines. PRCC encodes proline-rich protein PRCC (ppPRCC) of unknown function associated with papillary renal cell carcinoma. HPD encodes 4-hydroxyphenylpyruvate dioxygenase (HPPD), a key enzyme in tyrosine degradation whose Df causes autosomal recessive Tyrosinemia type 3, characterized by mental retardation. Interestingly, lines with a partial Df of HPD ortholog showed increased intraneuronal accumulation of Aβ42 that coincided with geotaxis impairment. These previously undetected modifiers of Aβ42 neurotoxicity in Drosophila warrant further study to validate their possible role and significance in the pathogenesis of sporadic AD.
Keywords: Alzheimer’s disease; Drosophila; amyloid β; dementia; genetic screen; neurodegeneration.
Figures

References
-
- Aksenov M. Y., Aksenova M. V., Markesbery W. R., Butterfield D. A. (1998). Amyloid β-peptide (1–40)-mediated oxidative stress in cultured hippocampal neurons. Protein carbonyl formation, CK BB expression and the level of Cu, Zn and Mn SOD mRNA. J. Mol. Neurosci. 10, 181–192. 10.1007/bf02761773 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

