The double-edged sword of (re)expression of genes by hypomethylating agents: from viral mimicry to exploitation as priming agents for targeted immune checkpoint modulation
- PMID: 28359286
- PMCID: PMC5374693
- DOI: 10.1186/s12964-017-0168-z
The double-edged sword of (re)expression of genes by hypomethylating agents: from viral mimicry to exploitation as priming agents for targeted immune checkpoint modulation
Abstract
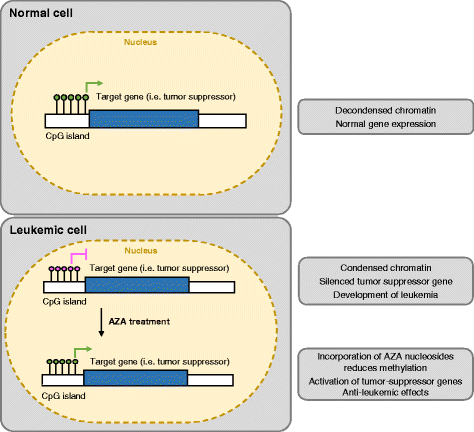
Hypomethylating agents (HMAs) have been widely used over the last decade, approved for use in myelodysplastic syndrome (MDS), chronic myelomonocytic leukemia (CMML) and acute myeloid leukemia (AML). The proposed central mechanism of action of HMAs, is the reversal of aberrant methylation in tumor cells, thus reactivating CpG-island promoters and leading to (re)expression of tumor suppressor genes. Recent investigations into the mode of action of azacitidine (AZA) and decitabine (DAC) have revealed new molecular mechanisms that impinge on tumor immunity via induction of an interferon response, through activation of endogenous retroviral elements (ERVs) that are normally epigenetically silenced. Although the global demethylation of DNA by HMAs can induce anti-tumor effects, it can also upregulate the expression of inhibitory immune checkpoint receptors and their ligands, resulting in secondary resistance to HMAs. Recent studies have, however, suggested that this could be exploited to prime or (re)sensitize tumors to immune checkpoint inhibitor therapies. In recent years, immune checkpoints have been targeted by novel therapies, with the aim of (re)activating the host immune system to specifically eliminate malignant cells. Antibodies blocking checkpoint receptors have been FDA-approved for some solid tumors and a plethora of clinical trials testing these and other checkpoint inhibitors are under way. This review will discuss AZA and DAC novel mechanisms of action resulting from the re-expression of pathologically hypermethylated promoters of gene sets that are related to interferon signaling, antigen presentation and inflammation. We also review new insights into the molecular mechanisms of action of transient, low-dose HMAs on various tumor types and discuss the potential of new treatment options and combinations.
Keywords: Acute myeloid leukemia; Azacitidine; Cancer; DNA methylation; Decitabine; Endogenous retroviral elements; Hypomethylating agents; Immune checkpoint blockade; Immune checkpoint inhibitors; Tumor microenvironment.
Figures



Similar articles
-
Digging deep into "dirty" drugs - modulation of the methylation machinery.Drug Metab Rev. 2015 May;47(2):252-79. doi: 10.3109/03602532.2014.995379. Epub 2015 Jan 8. Drug Metab Rev. 2015. PMID: 25566693 Free PMC article. Review.
-
Hypomethylating agents in combination with immune checkpoint inhibitors in acute myeloid leukemia and myelodysplastic syndromes.Leukemia. 2018 May;32(5):1094-1105. doi: 10.1038/s41375-018-0070-8. Epub 2018 Feb 22. Leukemia. 2018. PMID: 29487386 Free PMC article. Review.
-
Hypomethylating Agents and Immunotherapy: Therapeutic Synergism in Acute Myeloid Leukemia and Myelodysplastic Syndromes.Front Oncol. 2021 Feb 25;11:624742. doi: 10.3389/fonc.2021.624742. eCollection 2021. Front Oncol. 2021. PMID: 33718188 Free PMC article. Review.
-
Hypomethylation and up-regulation of PD-1 in T cells by azacytidine in MDS/AML patients: A rationale for combined targeting of PD-1 and DNA methylation.Oncotarget. 2015 Apr 20;6(11):9612-26. doi: 10.18632/oncotarget.3324. Oncotarget. 2015. PMID: 25823822 Free PMC article.
-
Decitabine has a biphasic effect on natural killer cell viability, phenotype, and function under proliferative conditions.Mol Immunol. 2013 Jul;54(3-4):296-301. doi: 10.1016/j.molimm.2012.12.012. Epub 2013 Jan 16. Mol Immunol. 2013. PMID: 23328088
Cited by
-
Epigenetic treatment-mediated modulation of PD-L1 predicts potential therapy resistance over response markers in myeloid malignancies: A molecular mechanism involving effectors of PD-L1 reverse signaling.Oncol Lett. 2019 Feb;17(2):2543-2550. doi: 10.3892/ol.2018.9841. Epub 2018 Dec 17. Oncol Lett. 2019. PMID: 30675316 Free PMC article.
-
Decitabine enhances targeting of AML cells by CD34+ progenitor-derived NK cells in NOD/SCID/IL2Rgnull mice.Blood. 2018 Jan 11;131(2):202-214. doi: 10.1182/blood-2017-06-790204. Epub 2017 Nov 14. Blood. 2018. PMID: 29138222 Free PMC article.
-
Genomic Study of COVID-19 Corona Virus Excludes Its Origin from Recombination or Characterized Biological Sources and Suggests a Role for HERVS in Its Wide Range Symptoms.Cytol Genet. 2020;54(6):588-604. doi: 10.3103/S0095452720060031. Epub 2021 Jan 15. Cytol Genet. 2020. PMID: 33487779 Free PMC article.
-
Selected nucleos(t)ide-based prescribed drugs and their multi-target activity.Eur J Pharmacol. 2019 Dec 15;865:172747. doi: 10.1016/j.ejphar.2019.172747. Epub 2019 Oct 18. Eur J Pharmacol. 2019. PMID: 31634460 Free PMC article. Review.
-
Activating PAX gene family paralogs to complement PAX5 leukemia driver mutations.PLoS Genet. 2018 Sep 14;14(9):e1007642. doi: 10.1371/journal.pgen.1007642. eCollection 2018 Sep. PLoS Genet. 2018. PMID: 30216339 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

