Prokaryote genome fluidity is dependent on effective population size
- PMID: 28362722
- PMCID: PMC5520154
- DOI: 10.1038/ismej.2017.36
Prokaryote genome fluidity is dependent on effective population size
Abstract
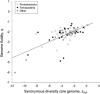
Many prokaryote species are known to have fluid genomes, with different strains varying markedly in accessory gene content through the combined action of gene loss, gene gain via lateral transfer, as well as gene duplication. However, the evolutionary forces determining genome fluidity are not yet well understood. We here for the first time systematically analyse the degree to which this distinctive genomic feature differs between bacterial species. We find that genome fluidity is positively correlated with synonymous nucleotide diversity of the core genome, a measure of effective population size Ne. No effects of genome size, phylogeny or homologous recombination rate on genome fluidity were found. Our findings are consistent with a scenario where accessory gene content turnover is for a large part dictated by neutral evolution.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
-
- Gogarten JP, Doolittle WF, Lawrence JG. (2002). Prokaryotic evolution in light of gene transfer. Mol Biol Evol 19: 2226–2238. - PubMed
-
- Gogarten JP, Townsend JP. (2005). Horizontal gene transfer, genome innovation and evolution. Nat Rev Microbiol 3: 679–687. - PubMed
-
- Kimura M (1984). The Neutral Theory of Molecular Evolution. Cambridge University Press: Cambridge, UK..
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources