Evolved genetic and phenotypic differences due to mitochondrial-nuclear interactions
- PMID: 28362806
- PMCID: PMC5375140
- DOI: 10.1371/journal.pgen.1006517
Evolved genetic and phenotypic differences due to mitochondrial-nuclear interactions
Abstract
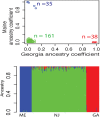
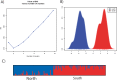
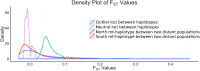
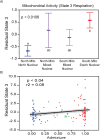
The oxidative phosphorylation (OxPhos) pathway is responsible for most aerobic ATP production and is the only pathway with both nuclear and mitochondrial encoded proteins. The importance of the interactions between these two genomes has recently received more attention because of their potential evolutionary effects and how they may affect human health and disease. In many different organisms, healthy nuclear and mitochondrial genome hybrids between species or among distant populations within a species affect fitness and OxPhos functions. However, what is less understood is whether these interactions impact individuals within a single natural population. The significance of this impact depends on the strength of selection for mito-nuclear interactions. We examined whether mito-nuclear interactions alter allele frequencies for ~11,000 nuclear SNPs within a single, natural Fundulus heteroclitus population containing two divergent mitochondrial haplotypes (mt-haplotypes). Between the two mt-haplotypes, there are significant nuclear allele frequency differences for 349 SNPs with a p-value of 1% (236 with 10% FDR). Unlike the rest of the genome, these 349 outlier SNPs form two groups associated with each mt-haplotype, with a minority of individuals having mixed ancestry. We use this mixed ancestry in combination with mt-haplotype as a polygenic factor to explain a significant fraction of the individual OxPhos variation. These data suggest that mito-nuclear interactions affect cardiac OxPhos function. The 349 outlier SNPs occur in genes involved in regulating metabolic processes but are not directly associated with the 79 nuclear OxPhos proteins. Therefore, we postulate that the evolution of mito-nuclear interactions affects OxPhos function by acting upstream of OxPhos.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Comment in
-
Fishing for adaptive epistasis using mitonuclear interactions.PLoS Genet. 2017 Mar 31;13(3):e1006662. doi: 10.1371/journal.pgen.1006662. eCollection 2017 Mar. PLoS Genet. 2017. PMID: 28362804 Free PMC article. No abstract available.
Similar articles
-
Gene by environmental interactions affecting oxidative phosphorylation and thermal sensitivity.Am J Physiol Regul Integr Comp Physiol. 2016 Jul 1;311(1):R157-65. doi: 10.1152/ajpregu.00008.2016. Epub 2016 May 25. Am J Physiol Regul Integr Comp Physiol. 2016. PMID: 27225945
-
Mitochondrial-nuclear interactions: compensatory evolution or variable functional constraint among vertebrate oxidative phosphorylation genes?Genome Biol Evol. 2013;5(10):1781-91. doi: 10.1093/gbe/evt129. Genome Biol Evol. 2013. PMID: 23995460 Free PMC article.
-
Evolution of nuclearly encoded mitochondrial genes in Metazoa.Gene. 2005 Jul 18;354:181-8. doi: 10.1016/j.gene.2005.03.046. Gene. 2005. PMID: 15975737
-
Not all mitochondrial DNAs are made equal and the nucleus knows it.IUBMB Life. 2021 Mar;73(3):511-529. doi: 10.1002/iub.2434. Epub 2020 Dec 25. IUBMB Life. 2021. PMID: 33369015 Free PMC article. Review.
-
Metazoan OXPHOS gene families: evolutionary forces at the level of mitochondrial and nuclear genomes.Biochim Biophys Acta. 2006 Sep-Oct;1757(9-10):1171-8. doi: 10.1016/j.bbabio.2006.04.021. Epub 2006 May 4. Biochim Biophys Acta. 2006. PMID: 16781661 Review.
Cited by
-
The Role of Mitonuclear Incompatibility in Bipolar Disorder Susceptibility and Resilience Against Environmental Stressors.Front Genet. 2021 Mar 16;12:636294. doi: 10.3389/fgene.2021.636294. eCollection 2021. Front Genet. 2021. PMID: 33815470 Free PMC article. Review.
-
Mitonuclear mismatch alters nuclear gene expression in naturally introgressed Rhinolophus bats.Front Zool. 2021 Sep 6;18(1):42. doi: 10.1186/s12983-021-00424-x. Front Zool. 2021. PMID: 34488775 Free PMC article.
-
Fishing for adaptive epistasis using mitonuclear interactions.PLoS Genet. 2017 Mar 31;13(3):e1006662. doi: 10.1371/journal.pgen.1006662. eCollection 2017 Mar. PLoS Genet. 2017. PMID: 28362804 Free PMC article. No abstract available.
-
Mitonuclear interactions shape both direct and parental effects of diet on fitness and involve a SNP in mitoribosomal 16s rRNA.PLoS Biol. 2023 Aug 21;21(8):e3002218. doi: 10.1371/journal.pbio.3002218. eCollection 2023 Aug. PLoS Biol. 2023. PMID: 37603597 Free PMC article.
-
Transcriptomic analysis provides insights into molecular mechanisms of thermal physiology.BMC Genomics. 2022 Jun 4;23(1):421. doi: 10.1186/s12864-022-08653-y. BMC Genomics. 2022. PMID: 35659182 Free PMC article.
References
-
- Chinnery PF. New approaches to the treatment of mitochondrial disorders. Reprod Biomed Online. 2004;8(1):16–23. - PubMed
-
- Dowling DK, Friberg U, Hailer F, Arnqvist G. Intergenomic epistasis for fitness: within-population interactions between cytoplasmic and nuclear genes in Drosophila melanogaster. Genetics. 2007;175(1):235–44. Epub 2006/12/08. PubMed Central PMCID: PMCPMC1774999. 10.1534/genetics.105.052050 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources

