A furoviral replicase recruits host HSP70 to membranes for viral RNA replication
- PMID: 28367995
- PMCID: PMC5377427
- DOI: 10.1038/srep45590
A furoviral replicase recruits host HSP70 to membranes for viral RNA replication
Abstract
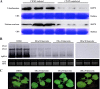
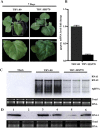
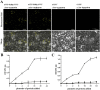
Many host factors have been identified to be involved in viral infection. However, although furoviruses cause important diseases of cereals worldwide, no host factors have yet been identified that interact with furoviral genes or participate in the viral infection cycle. In this study, both TaHSP70 and NbHSP70 were up-regulated in Chinese wheat mosaic furovirus (CWMV)-infected plants. Their overexpression and inhibition were correlated with the accumulation of viral genomic RNAs, suggesting that the HSP70 genes could be necessary for CWMV infection. The subcellular distributions of TaHSP70 and NbHSP70 were significantly affected by CWMV infection or by infiltration of RNA1 alone. Further assays showed that the viral replicase encoded by CWMV RNA1 interacts with both TaHSP70 and NbHSP70 in vivo and vitro and that its region aa167-333 was responsible for the interaction. Subcellular assays showed that the viral replicase could recruit both TaHSP70 and NbHSP70 from the cytoplasm or nucleus to the granular aggregations or inclusion-like structures on the intracellular membrane system, suggesting that both HSP70s may be recruited into the viral replication complex (VRC) to promote furoviral replication. This is the first host factor identified to be involved in furoviral infection, which extends the list and functional scope of HSP70 chaperones.
Conflict of interest statement
The authors declare no competing financial interests.
Figures






Similar articles
-
Root-specific role for Nicotiana benthamiana RDR6 in the inhibition of Chinese wheat mosaic virus accumulation at higher temperatures.Mol Plant Microbe Interact. 2013 Oct;26(10):1165-75. doi: 10.1094/MPMI-05-13-0137-R. Mol Plant Microbe Interact. 2013. PMID: 23777430
-
Heat shock protein 70 is necessary for Rice stripe virus infection in plants.Mol Plant Pathol. 2014 Dec;15(9):907-17. doi: 10.1111/mpp.12153. Epub 2014 Jun 30. Mol Plant Pathol. 2014. PMID: 24823923 Free PMC article.
-
Analysis of small RNAs derived from Chinese wheat mosaic virus.Arch Virol. 2014 Nov;159(11):3077-82. doi: 10.1007/s00705-014-2155-7. Epub 2014 Jul 6. Arch Virol. 2014. PMID: 24997977
-
Coat Protein Regulation by CK2, CPIP, HSP70, and CHIP Is Required for Potato Virus A Replication and Coat Protein Accumulation.J Virol. 2017 Jan 18;91(3):e01316-16. doi: 10.1128/JVI.01316-16. Print 2017 Feb 1. J Virol. 2017. PMID: 27852853 Free PMC article.
-
Viral sensing of the subcellular environment regulates the assembly of new viral replicase complexes during the course of infection.J Virol. 2015 May;89(10):5196-9. doi: 10.1128/JVI.02973-14. Epub 2015 Mar 4. J Virol. 2015. PMID: 25741009 Free PMC article. Review.
Cited by
-
Hsc70-2 is required for Beet black scorch virus infection through interaction with replication and capsid proteins.Sci Rep. 2018 Mar 14;8(1):4526. doi: 10.1038/s41598-018-22778-9. Sci Rep. 2018. PMID: 29540800 Free PMC article.
-
Regulation of heat shock proteins 70 and their role in plant immunity.J Exp Bot. 2022 Apr 5;73(7):1894-1909. doi: 10.1093/jxb/erab549. J Exp Bot. 2022. PMID: 35022724 Free PMC article. Review.
-
Oligomeric SWEET1g of Solanum tuberosum confers resistance to potato virus Y and Potato virus X.Plant J. 2025 Aug;123(3):e70400. doi: 10.1111/tpj.70400. Plant J. 2025. PMID: 40758984 Free PMC article.
-
Stvb-i, a Rice Gene Conferring Durable Resistance to Rice stripe virus, Protects Plant Growth From Heat Stress.Front Plant Sci. 2020 May 8;11:519. doi: 10.3389/fpls.2020.00519. eCollection 2020. Front Plant Sci. 2020. PMID: 32457773 Free PMC article.
-
Plasmodesmal endoplasmic reticulum proteins regulate intercellular trafficking of cucumber mosaic virus in Arabidopsis.J Exp Bot. 2023 Aug 17;74(15):4401-4414. doi: 10.1093/jxb/erad190. J Exp Bot. 2023. PMID: 37210666 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

