A genome-wide study of Hardy-Weinberg equilibrium with next generation sequence data
- PMID: 28374190
- PMCID: PMC5429372
- DOI: 10.1007/s00439-017-1786-7
A genome-wide study of Hardy-Weinberg equilibrium with next generation sequence data
Abstract
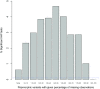
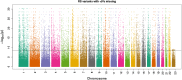
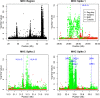
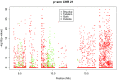
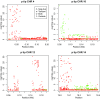
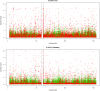
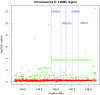
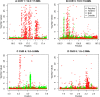
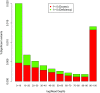
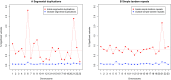
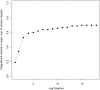
Statistical tests for Hardy-Weinberg equilibrium have been an important tool for detecting genotyping errors in the past, and remain important in the quality control of next generation sequence data. In this paper, we analyze complete chromosomes of the 1000 genomes project by using exact test procedures for autosomal and X-chromosomal variants. We find that the rate of disequilibrium largely exceeds what might be expected by chance alone for all chromosomes. Observed disequilibrium is, in about 60% of the cases, due to heterozygote excess. We suggest that most excess disequilibrium can be explained by sequencing problems, and hypothesize mechanisms that can explain exceptional heterozygosities. We report higher rates of disequilibrium for the MHC region on chromosome 6, regions flanking centromeres and p-arms of acrocentric chromosomes. We also detected long-range haplotypes and areas with incidental high disequilibrium. We report disequilibrium to be related to read depth, with variants having extreme read depths being more likely to be out of equilibrium. Disequilibrium rates were found to be 11 times higher in segmental duplications and simple tandem repeat regions. The variants with significant disequilibrium are seen to be concentrated in these areas. For next generation sequence data, Hardy-Weinberg disequilibrium seems to be a major indicator for copy number variation.
Conflict of interest statement
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Figures

References
-
- Crow JF, Kimura M. An introduction to population genetics theory. New York: Harper & Row Publishers; 1970.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

